RESEARCH ARTICLE
Uncovering the anti-bacterial potential of wildly growing Chamaedorea seifrizii fruits targeting peptidoglycan editing factor proteins: chemical profiling, in-silico analysis and wet lab validation
Sciences of Phytochemistry|Vol. 5, Issue 1, pp. 135-146 (2026)
Received
Sep 12, 2025Revised
Feb 5, 2026Accepted
Feb 12, 2026Published
May 7, 2026
Abstract
Introduction
Multidrug-resistant (MDR) bacteria have emerged as one of the biggest threats to human health as result of the overuse and abuse of antibiotics in global healthcare (1). Medicinal plants have been the main source of therapeutic remedies from ancient times, despite their crucial significance in modern medicine. Phytochemicals, which are produced from plants, have strong effects when used alone or in conjunction with traditional antibiotics. Plant extracts, rich in polyphenols and tannins, offer a promising alternative to combat antibiotic-resistant bacteria, often by enhancing the efficacy of conventional antibiotics or acting as efflux pump inhibitors (1). Bacteria employ a unique enzyme known as Peptidoglycan Editing Factor (PdeF) to identify and eliminate the offending amino acids in order to stop this. It is a bacterial cytoplasmic amidase that serves as a peptidoglycan (PG) biosynthesis quality control mechanism (2). Cell integrity and shape depend on PG, a characteristic of the bacterial cell membrane Understanding the cell wall biosynthesis pathway aids in the development of novel tactics to stop bacterial growth. Therefore, PG, is a key target for in silico based studies. It controls precursor-editing process that preserves the integrity of the peptidoglycan structure that is crucial element of bacterial survival. Therefore, PdeF is essential for preserving the bacterial cell wall's mechanical and structural integrity, which is necessary for defense against osmotic pressure (3). Importantly, PdeF represents a relatively underexplored target within the peptidoglycan biosynthesis pathway compared to classical targets such as transpeptidases targeted by β-lactam antibiotics. Unlike conventional targets, PdeF is involved in the proofreading and editing of peptidoglycan precursors, a critical step for maintaining cell wall fidelity. Disruption of this process may lead to accumulation of defective precursors and compromised cell wall integrity, making PdeF a promising alternative antibacterial target, particularly in the context of multidrug resistance.
Materials and Methods
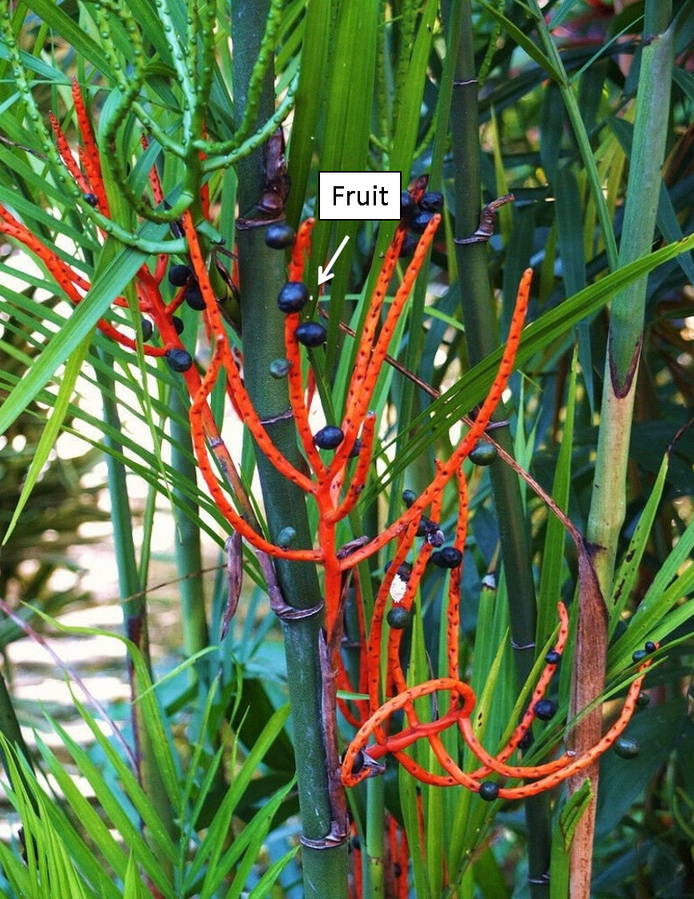
Plant collection Wildly growing C. seifrizii Figure 1 plant samples (fruits) were collected from garden area close to study area (31.30 latitudes North and 75.60 longitude East, Jalandhar, Punjab, India). The botany department confirmed the plant, and BT106 was entered on the voucher and deposited in biotech herbarium. The climate of this city is humid subtropical, with long, hot summers and cold winters. The experiment was conducted in January – April 2025.

Preparation of Extracts
After rinsing the gathered fruits with distilled water to get rid of any sticking material, they were shade-dried for 7 days at room temperature (25–28 °C) until their weight remained consistent. A mortar and pestle were used to further decrease the size in powder form. After that, it was sieved, processed using an electrical grinder, and kept at 4 °C for further examination. Phytocompounds from powdered C. seifrizii fruits (3 g) were extracted using methanol/water solvent (8: 2 v/v). The mixture was manually mixed by shaking, vortexed, and was incubated overnight in orbital shaker at ambient temperature prior to being spun down for fifteen min at 10, 000 rpm and 4 °C. The resulting extract was collected, filtered using Whatman No. 1 filter paper the filtrate was concentrated using a rotary evaporator at 64.5 °C (% yield: 5.7%) and stored at 4 °C for further examination. Extracts were designated as C. seifrizii fruit methanol extract: CFME.
GC-FID Analysis of Extracts
According to Amrita et al. , (9), a gas chromatography flame ionization detector (GC-FID) system (Chemtron Pvt ltd. 2045) was used to identify phytochemicals in methanol extracts of fruits of C. seifrizii. Sample volume of 2 µL was used for analysis in a split mode. 10% OV-17 was placed onto 80–100% mesh Chromosorb W (HP) in a 2-meter stainless-steel column that was part of the GC apparatus. 30 mL per min of nitrogen gas (99.9995% purity) was employed as the gas used for the mobile phase. The temperature of the injector and detector was retained at 200 °C and 250 °C. The oven's ramping settings were 110 °C at first, then 200 °C at a rate of 2 °C per min. By comparing the relative retention time with authenticated standards and data from the literature, bioactive chemicals were found (10).
Fingerprint Analysis of Extracts
Understanding the chemical characteristics of the numerous phytochemical substances and functional groups found in CFME is crucial for understanding their therapeutic effects. Thus in order for functional characterization, it was subjected to fingerprint analysis using ultraviolet spectroscopy (UV), Fourier Transform Infrared Spectroscopy (FT-IR) and fluorescence spectroscopy (9). For UV, 1 mL of CFME was subjected to scan from 200 to 700 nm and peaks were recorded on UV-VIS spectrophotometer (Labtronics, India). To identify functional groups in extracts' active components, FT-IR analysis was done. About 10 µL of extracts were used for this little quantity and were scanned in the 400–4000 cm-1 range using an FT-IR spectrophotometer (Perkin Elmer, USA). For fluorescence examination, 3 mL of CFME was used and peaks were recorded (Perkin Elmer, USA spectrophotometer). The NN07 method was applied, with a 10nm slit width, emission ranging from 410 nm - 700 nm, and a scan speed of 240nm/min at an excitation wavelength of 380 nm. Experiments were carried out at room temperature (25 °C - 30 °C).
In-vitro Evaluation of Antibacterial Activity
Using the agar disc diffusion technique (9), the antibacterial activity of CFME was evaluated against four test organisms: Pseudomonas aeruginosa (MTCC 424), G+ g-positive Staphylococcus aureus (MTCC 3160), Bacillus subtilis (MTCC 121), and G-Escherichia coli (MTCC 40). Sterilized paper discs with a diameter of 5 mm were impregnated with CFME (80 µg). For every microbial strain, 12-h-old cultures were used to establish new inoculums. The LB-Agar plates were swabbed with bacterial suspension and then left to dry for half an h. The paper discs loaded with CFME were then placed in the center of the petri plates. The plates were incubated for 24 h at 37 °C after being left at room temperature for 20 min. Gentamycin was used as the positive control (100 µg/disc). Methanol was used negative control. Using a trans-illuminator, one can see the zone of inhibition (ZOI: mm) surrounding the disc after incubation; this radius is indicative of the antibacterial activity of extracts.
Membrane Integrity Assay
The effect of CFME on membrane integrity was examined against the bacterial strains P. aeruginosa (MTCC 424, G-g-negative), E. coli (MTCC 40 G-g-negative), S. aureus (MTCC 3160 G+g-positive), and B. subtilis (MTCC 121 G+g-positive) in accordance with the protocol described by (Nirmala et al. , 2020) with some modifications. CFME (80 µL) was used to treat the bacteria (1x108 CFU/mL). Bacterial cells were treated with 200 µL of Triton X-100 as a positive control. As a negative control, untreated cells were used. After being incubated in an orbital shaker incubator at 37 °C and 150 rpm for 12 h, the treated cells were monitored. One mL of the sample of each bacterial strain was extracted from each flask after an equivalent amount of time had elapsed. The cytoplasmic contents were then extracted by centrifuging the cells for 10 min at 6000 rpm. The resultant supernatant was collected and analyzed at 260 nm using a Labtronics single beam UV spectrophotometer.
Computational Studies
To validate the wet lab study, the in-silico analysis was conducted using major phytocompounds osthole against targeting PdeF, associated proteins involved in the bacterial cell wall synthesis.
Preparation of Ligands
Major abundant bioactive compound viz: osthole in CFME was chosen as ligand for molecular docking studies. Ligands were prepared by retrieving SMILES from the NCBI-PubChem database and 3-D structures were generated using UCSF-chimera. Gentamycin drug was used as a standard inhibitor.
Target Protein Preparation
The crystal structures PdeF (pdb id: 7fbg) was employed as target enzymes in this study. PdeF structure was obtained from the RCSB-PDB database. Target enzyme receptors were prepared for docking studies using the dock prep set up in the Chimera software. Dock prep is an optimization part that corrects structure, charge anomalies, and atomic bond length.
Molecular Docking
To investigate the binding method of bioactive compound selected as ligand with members of PdeF protein, molecular docking was carried out by using Cb-dock2. Docking was executed by uploading the ligand and target enzyme molecules in a. pdb file to the cb-dock2 tool. The best-generated model in the. pdb file is downloaded and saved. The enzyme-ligand 2-D interactions were predicted by using the Biovia 2020 and UCSF Chimera tools.
Molecular Dynamic Simulation (MDS)
To further evaluate the binding stability, conformational behavior, and interaction dynamics between the top-ranked docked bioactive compounds (ligands) and the PdeF receptor, molecular dynamics simulation (MDS) was carried out following the protocol described by Chauhan and Sharma (2025). The simulations were performed using GROMACS version 2019.2 to analyze the physicochemical behavior of the selected ligand–PdeF complexes under dynamic conditions. Initially, energy minimization was conducted using the steepest descent algorithm for 5000 steps to eliminate steric clashes and reduce unfavorable contacts within the system. The minimized complex was then placed in a cubic periodic simulation box with a distance of 0.5 nm from the box edges and solvated using the simple point charge (SPC) water model. To ensure system neutrality and physiological relevance, appropriate numbers of Na⁺ and Cl⁻ counter ions were added to achieve a salt concentration of 0.15 M. Subsequently, the system underwent equilibration under NPT (constant number of particles, pressure, and temperature) ensemble conditions, followed by a production run of 50 ns. The stability and structural fluctuations of the complexes were evaluated using parameters such as root mean square deviation (RMSD), root mean square fluctuation (RMSF), and radius of gyration (Rg). In addition, the binding free energy of the complexes was calculated using the molecular mechanics generalized Born surface area (MM-GBSA) approach, based on equilibrated trajectories obtained from the GROMACS simulations, to quantify the strength and stability of ligand–receptor interactions.
Drug-likeness and ADMET
Physiochemical properties and Drug-likeness properties of ligand (osthole) were conducted by employing SWISSADME. Pharmacokinetics ADMET (Absorption, distribution, Metabolism, excretion and Toxicity), studies were conducted using ProTox 3.0 tools and pkCSM tools.
Statistical Analysis
Microsoft Excel and SPSS 26.0 of the Statistical Product and Service Solution (SPSS) were utilized to analyze the collected data of ZOI. To determine the statistical significance of the data between groups, Tukey HSD post-hoc statistical analysis was conducted. At p < 0.05, mean differences were deemed significant. The trials were conducted in triplicate (n = 3), and the results were presented as Mean ± Standard Deviation.
Results and Discussion
Fingerprint Analysis of C. seifrizii Fruit (cfme) Extracts
The existence of phytochemicals was shown by the qualitative UV spectroscopy profile of CFME, which showed several strong peak from at about 350 nm with absorbance of 1.3 Figure 2A. Early study by Sowmya et al (11) reported that certain kinds of phytosubstances can be identified by the presence of peaks in particular wavelength bands. For instance, the absorption in the 200–400 nm range points to heteroatoms and unsaturated groups, which are characteristics of organic chromophores. The term "UV profile" describes a phytochemical's distinct ultraviolet-visible (UV-Vis) spectroscopy signature, which shows particular absorption bands that signify the presence of phytochemicals (12). The presence polyphenolics was further validated by quantitative analysis test which revealed contents is a following order: Phenolics: 392.23 ± 0.23 [µgGAE gDW-1], Flavonoids: 290.33 ± 3.85 [µgRUT gDW-1], Tannins: 208.85 ± 2.12 [µgAA gDW-1] Supplementary Table 1.
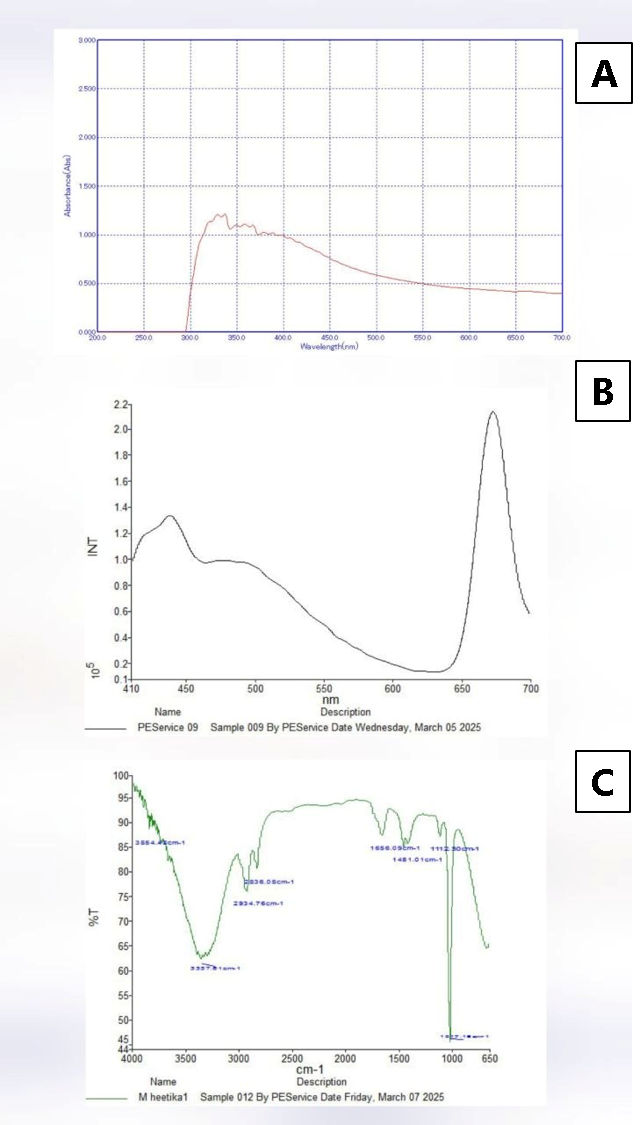
Fluorescent spectroscopy is used to create a "fingerprint" for detecting compounds (13). This method relies on the unique spectral properties of compounds, allowing for rapid, non-destructive and sensitive analysis (14). The fluorescence emission spectrum of CFME is shown in Figure 2B. One prominent peak lambda at 450 nm (green fluorescent region/) was seen in CFME, suggesting the presence of secondary metabolites compounds in CFME. Flavonoids, terpenoids, and flavins may be the biomolecules found in the green fluorescent area (14). Donaldson, (15) reported that flavonoids are of interest as potential therapeutic agents and are autofluorescent with emission at green, yellow and orange wavelengths in plants such as Eysenhardtia polystachya. Singh and Mishra (17) also reported the presence of polyphenolics and anthocyanins in vegetables extracts in the fluorescent range of 500-650 nm. A wide band in the 650 nm (red fluorescent region) is frequently ascribed to alkaloids, polyacetylenes chemicals and products of the Maillard process when stimulated at 280 nm (14, 15). Plant’s fluorescence emission spectrum may be thought of as its chemical signature, reflecting how it reacts to biotic and abiotic environmental challenges and highlighting the importance of secondary metabolites. When exposed to radiation of an appropriate wavelength, a variety of metabolites, including proteins, coenzymes, flavonoids, phenolics, alkaloids, chlorophylls, and components of cell walls, exhibit autofluorescent behavior (14). The accumulation of fluorescent secondary metabolites, such as terpenoids, carotenoids, and phenolics, which are critical for the proper growth and development of plants, is thought to shield cellular structures from the damaging effects of abiotic stressors. Szukay et al. (14) studied bee products and cited that the presence of a distinct mix of vitamins, phenolic compounds, aromatic amino acids, and Maillard reaction products gives fluorescent characteristics in bee products. Bui et al. (15) reported that many chemical compounds such as polyphenols and flavonoids are fluorophores in spice extracts.
FT-IR fingerprint analysis of extracts provides a rapid, non-destructive, and low-cost method to identify chemical composition by detecting functional groups and vibrational signatures, particularly in the 1500–500 cm-1. The spectrum, often viewed in the fingerprint region (500– 1500 cm -1), acts as a unique signature of the compound's structure, allowing for distinguishing between species or plant parts. FT-IR profile of CFME is displayed in Figure 2C. Four major sections may be distinguished in the FT-IR spectrum: the first is 4000-2500 cm-1, the s is 2500-2000 cm-1, the third is 2000-1500 cm-1, and the fourth is 1500-400 cm-1. Due to its abundance of complex peaks, the fourth area of the infrared spectrum is often referred to as the fingerprint region (16). FT-IR spectrum was measured at the range of 4000-650 cm-1 that showed different peaks. Supplementary Table 1 provides comprehensive details on every peak. The FT-IR peaks showed variance across in extract. The FT-IR spectra of both CFME showed a wide band in the 3400–3200 cm-1 region, which was caused by the OH– group and therefore indicated the presence of phenolic chemicals. This region exhibits both symmetric (sym) and asymmetric (asym) stretching due to polymeric hydroxyl group (O–H) and H-bonded stretching that is typical of polyphenolic compounds (17). Other distinct bands were seen in CFME extract at around 2800–2900 cm-1, which was caused by C–H stretching from the alkanes. This was due to the -CH, -CH2, and -CH3 stretching vibrations, which are produced from the carbohydrates and sugars in herbal extracts (17). The presence of aromatic chemicals is strongly correlated with these group frequencies. The 1500–400 cm-1 area of the FT-IR spectrum, showed variations. The area, which stretched between 1500 and 400 cm-1, is sometimes referred to as the fingerprint region due to the abundance of distinctive single bands with low intensities that are linked to particular functional groups. C–H, C–O, C–N, and P–O bonds are among these groupings. It is the fourth main area in the infrared spectrum, so named because it has a great deal of complicated peaks. Fourier Transform Infrared Spectroscopy (FT-IR), often referred to as “fingerprint" method because it produce distinct "fingerprint," or spectral pattern which is particular to a molecule or a compound. This unique fingerprint pattern enables the identification and examination of chemical compositions and structures (18). As per earlier research, herbal extracts may be identified and characterized using FT-IR spectroscopy, which also helps with quality control by identifying functional groups. It offers a non-destructive technique for sample analysis that calls for little setup and little amounts. Numerous researches have been conducted on the use of spectroscopic techniques, particularly Fourier-transform infrared spectroscopy (FT-IR), to identify bioactive chemicals in plants (19). These results imply that secondary metabolites are plentiful in C. seifrizii fruits (CFME) extracts. The existence of the aforementioned secondary metabolites may be the cause of C. seifrizii therapeutic qualities.
GC-FID Identification and Quantification of Bioactive Compounds Present in Chamaedorea Seifrizii Acetone Extracts
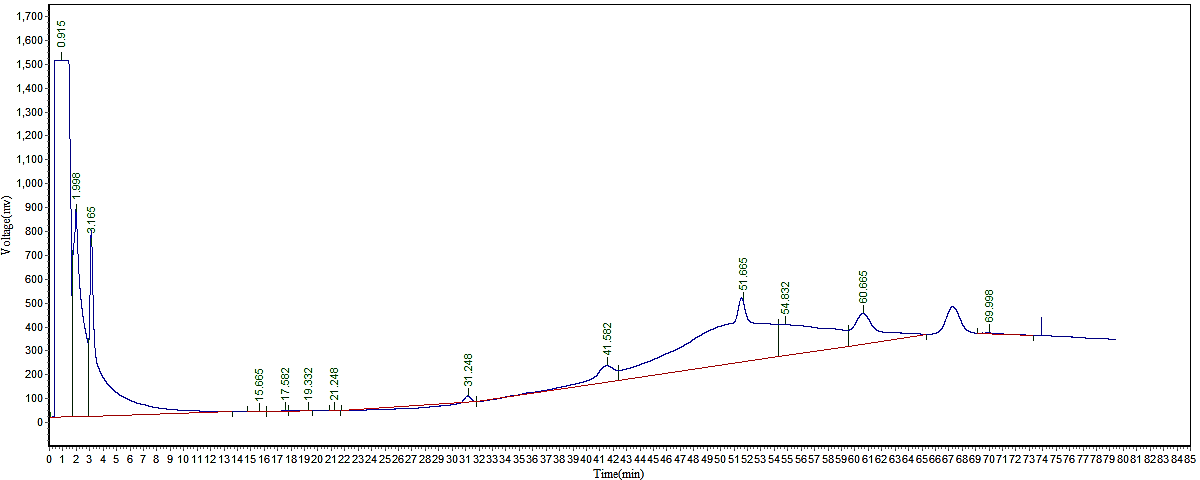
Figure 3 shows the typical GC-FID chromatogram, which show the peaks that correspond to several bioactive compounds found in the extracts of C. seifrizii methanol extract (CFME). It lists the compounds present in the CFME, together with their % (%) compositions, and retention time (RT). The GC–FID analysis of CFME led to the identification of 13 components. As shown in Figure 3, The dominant phytochemical compound was osthole, consistently quantified at 12.28% in the GC-FID analysis. These findings are similar to those of Duru, (20, 21), who used gas chromatography and a flame ionization detector to measure the phytochemical content in solvents and observed maximum amount in an ethanolic extract of Zea mays husk. Additionally, there have been reports of some of these components' biological activity. Numerous plants naturally contain osthole, which was identified as the major compound (12.28%) in the present study and resembles a coumarin derivative (22). According to certain research, osthole may have neuroprotective benefits (23). Based on this data, it was concluded that C. seifrizii is rich in phytochemical constituents that have a key biological activities, such as, antioxidant according to the reviewed literature. Consequently, the diverse range of physiological potentials and biological activities exhibited by C. seifrizii may be attributed to the existence of these bioactive chemicals.
In silico Analysis
Molecular Docking
PdeF (Peptidoglycan Editing Factor), which is necessary for the latter stages of peptidoglycan (PG) bacterial cell wall formation, was the focus of this in-silico investigation. Molecular docking is one of the efficient computational methods utilized in this scenario to forecast and detect the potential binding mode that takes place between a chemical and a certain target protein or receptor (27). Molecular docking is a robust and reliable technique for analyzing the binding affinity, pose orientation, and interaction pattern of small molecules in the active area of the target protein. The benefits of molecular docking for drug design and discovery are well established. In terms of time and resource investment, the dry lab approach offers a significant advantage over in vivo lab experiments (28). Molecular docking has been an essential part of in-silico drug research in recent years (28). This structure-based approach aims to predict the atomic-level interaction between a small chemical and a protein. This enables researchers to understand the fundamental biochemical process underlying this interaction and investigate how small molecules, such as bioactives, behave within the binding region of a target protein (29).
PdeF was used as target in this molecular docking study, with the most prevalent phytocompound osthole (12.28%), as consistently identified by GC-FID analysis of C. seifrizii methanol extracts (CFME). Table 1 shows the docking score for osthole bioactive molecule against target (PdeF) and other relevant information. Supplementary Figure 1 shows 3D model of PdeF complex with highly affinitized ligand osthole. Osthole demonstrated a binding energy of −6.8 kcal/mol, indicating a moderate interaction with the target protein.. Less binding energy indicates higher affinity of ligand towards target protein. When ligand-receptor interactions take place, binding affinity is a crucial element that must be taken into account. The ligand (compound) needs less energy to attach to the receptor if the binding affinity is high. Therefore, high binding affinity levels have a greater potential to interact with target macromolecules. Notably when positive drug gentamycin was docked to PdeF, binding energy was -6.7 kcal/mol. This suggests that osthole exhibits a moderate binding affinity towards PdeF protein, comparable to gentamycin; however, such docking scores should be interpreted cautiously as they represent predictive interactions rather than definitive binding confirmation. By examining the docked structures, significant interactions between the ligand and the protein are also found, such as alkyl-Pi-alkyl, and hydrogen bonding interactions. These interactions can provide information about the ligands' mechanism of action and guide further structural optimization (30). The ligand's capacity to form hydrophobic or hydrogen bonding contacts with the active site residues of the receptor protein during docking determines its affinity for the receptor. The two-dimensional interactions between osthole and the amino acid residues in the corresponding target protein's active region are depicted in Supplementary Figure 1.
Our results showed that osthole ligand could dock at the binding site of PdeF. Despite having somewhat different docking scores, pose orientations, and interaction patterns, both ligands preferentially occupied the binding region and formed polar and non-polar interactions including pi-pi stacking, alkyl, pi-alkyl, and hydrogen bonds (H-bond) in the PdeF’s enzyme's active sites via contact residues. For instance: it was observed that most stable complex osthole– PdeF was stabilized by 4 conventional H-bonds. The amino acids THR 24, LYS 267, GLN 15, which form conventional hydrogen bonds when the ligand binds to PdeF receptor, play an important role in antibacterial activity. This finding suggests a possible mechanism by which osthole may interact with PdeF; however, direct experimental validation is required to confirm this mode of action. In-depth analysis revealed that osthole showed affinity towards DUF152 domain of PdeF involving “A” chain residues similar to synthetic drug gentamycin. It is part of the core domain of PdeF proteins and is implicated in peptidoglycan editing (31).
Although bacterial peptidoglycan editing factors are acknowledged as a promising new target for antibacterial medicines, there are presently no licensed medications that particularly target them. Ongoing research is looking at novel enzymes in the larger peptidoglycan (PG) production pathway as possible pharmacological targets, although numerous current drug classes, including vancomycin, cephalosporins, and penicillins, target this process (32). Even though there aren't any particular medications available to block PdeF, the continued study in this field shows promise for the creation of new antibacterial treatments in the future that could circumvent existing resistance mechanisms (33). The results of the study suggest that C. seifrizii extracts could be the primary source of antibacterial substances. According to the data mentioned above, phytocompounds exhibited a high affinity for PdeF after ligand contact. Consequently, it was postulated that PdeF alteration results in a modification of the bacterial cell wall and enzyme conformation. By stopping bacterial viability, all of these occurrences lessen the bacteria's capacity to invade the host cell. Previous research has also shown through in-silico results that multi-pharmacological drugs have antibacterial properties that can regulate the proliferation of microbial strains. Osthole has potential antibacterial activity based on in silico molecular docking studies. Overall, these findings suggest that C. seifrizii-derived phytocompounds such as osthole may have potential as antibacterial candidates; however, further experimental validation is required to confirm these in silico predictions.
| Receptors | Ligand | Binding energy kcal/mol (Vina score) | Cavity volume (Å3) | Center (x, y, z) | Docking size (x, y, z) | Amino acids involved in interaction |
|---|---|---|---|---|---|---|
| PdeF: 7fbg | Osthole | -6.8 | 306 | -4, 19, 28 | 19, 19, 19 | Chain A: TYR13 LEU14 GLN15 TRP17 LYS18 GLU19 GLY21 ASN22 ILE23 THR24 ALA25 GLY26 LYS65 LEU66 GLN67 GLU114 TYR131 PRO133 GLY136 PHE266 LYS267 |
| PdeF: 7fbg | Gentamycin | -6.7 | 429 | 1, -5, 18 | 22, 22, 22 | Chain A: LYS30 ASP31 THR36 GLY37 SER38 PHE39 HIS40 ALA41 GLU244 GLN245 PHE247 PHE248 SER249 ARG252 ASP253 GLN254 LYS256 THR257 GLY258 |
Molecular Dynamics Studies
Best-docking scored model, osthole- PdeF complex, was chosen for a 50 ns all-atom MDS Molecular dynamics simulation (MDS) was used to verify docking results. MDS makes it easier to accurately analyze the dynamics of biological macromolecules in a controlled physiological setting. Using MDS, a computational method for analyzing the physical interactions in a biophysical system, the flexibility and structural alterations of docked complexes may be seen over the simulation time. According to Liu et al. (34), such an investigation would also evaluate the ligand's crucial binding interactions with important catalytic site residues and provide valuable insights into the dynamic behavior of the ligand and protein. Supplementary Figure 5 display the findings of the post-MD simulation studies, which is expressed as RMSD, RMSF and Rg as function of the structural stability.
In comparison to the reference beginning structure, RMSD calculates the average distance between atoms in a structure at a given point in the simulation. The total time-dependent structural divergence or similarity between the structures captured in the trajectory is therefore examined. Examining how RMSD values vary over time or under various circumstances might help one understand the complex's dynamics and stability (35). This graph displays the degree of ligand-protein binding site compatibility during the simulation. A falling pattern might indicate that the ligand is adopting a more advantageous binding position, whereas a rising pattern would indicate that it is exploring other orientations. A stable fit means that the ligand's binding form remains constant. Important information on the orientation and structure of the simulated ligands in relation to their binding pocket might be obtained by tracking RMSD trajectories. After a few nanoseconds, the target, achieved stable confirmation, as the plot of the osthole-PdeF complex demonstrates. The first binding to the substrate causes massive fluctuations between 0.5 nm and 1.0 nm in the RMSD trajectory upon ligand binding till 15 ns, followed by the equilibrium state at < 1.0 nm during simulation time. The ligand-protein's RMSD trajectory suggests that it could be trying to find a better binding posture during the simulation, based on the initial binding position. The trajectory seems to have established a better suitable binding position toward the conclusion of the experiment, stabilizing at around 0.9 nm until 50 ns. This finding proved that ligand binding stabilizes the protein structure. These results also validated docking studies, indicating the impactful role of C. seifrizii as antibacterial agent having therapeutic applications.
The macromolecular system's conformational stability may also be described using root-mean-square fluctuation (RMSF). It is an essential metric for assessing the stability and flexibility of complex systems during simulation. The protein's Cα atoms' RMSF values were computed and shown in relation to the residues Supplementary Figure 2. The complex's N-terminal (1–100 aa), middle part (100–300 aa), and C-terminal (300 aa–onwards) sections have been identified for investigation. With the exception of small peaks in the protein's C-terminal region, the amino acid residues in the osthole–PdeF complex under study showed very little fluctuation throughout the simulation and remained below 0.4 nm. These results showed that ligand binding had no appreciable effect on the protein's flexibility, with an overall RMSF tone ranging from 0.1 to 0.4 nm. Due to its loop shape, the protein's C-terminal displayed comparatively large RMSF variations.
The RMSF (Root Mean Square Fluctuation) values were found to be proportionately connected to the previous RMSD (Root Mean Square Deviation) trajectory analysis, providing robust evidence for the preferential stability of the osthole–PdeF complex. This correlation suggests that the binding of osthole effectively minimizes the local fluctuations of the protein residues, leading to a more rigid and stable conformation.
To further evaluate the structural integrity of the system, the Rg was investigated as a measure of protein compactness. As illustrated in Supplementary Figure 2, the Rg values of the protein backbone atoms were monitored over the course of the simulation. Following the initial equilibration phase, the Rg value for the osthole–PdeF complex decreased to 3.85 nm. This reduction indicates a more condensed and compact secondary structure upon ligand binding, reinforcing the findings from the RMSD and RMSF analyses regarding the system's overall rigidity.
Finally, the thermodynamic stability of the interaction was quantified using the MM-PBSA (Molecular Mechanics Poisson-Boltzmann Surface Area) method. This approach provides a comprehensive estimation of the binding energy by integrating molecular mechanics with continuum solvation models. The resulting binding free energy (ΔGbindΔGbind) for the osthole–PdeF complex was determined to be -65.2 ± 20.14 kJ mol⁻¹. This significant negative value confirms a high affinity and a spontaneous binding process, establishing osthole as a potent stabilizer for the PdeF protein.
Wet Lab Validation
Antibacterial Activity of Chamaedorea seifrizii Methanol Extract
In order to confirm the in-silico results, the wet lab experiment research was created to evaluate the antibacterial properties of CFME against four bacterial strains. The results (Supplementary Table 2 and Supplementary Figure 3) indicate that CFME exhibits mild to moderate antibacterial activity rather than strong inhibition. Upon CFME treatment, The observed zones of inhibition (ZOI), ranging from 0.2 cm to 1.7 cm, suggest limited antibacterial potency when compared to standard antibiotics. , with most predominant effect on P. aeruginosa (MTCC424) with ZOI 1.7 cm. S. aureus ((MTCC 3160) and B. subtilis (MTCC 121) exhibited moderate inhibitory activity with ZOI value 0.2 cm. Very interestingly, when compared with positive drug gentamycin, against bacterial strain MTCC 3160, notable ZOI were observed where as positive control was unable to inhibit the growth of the pathogen. Although antimicrobial activity of CFME may be attributed to the presence of the major phytocompound osthole (12.28%), identified consistently in GC-FID analysis. However, the antimicrobial role of other phytochemicals existing in minor amounts can’t be ruled out (36). C. seifrizii strong antibacterial qualities may be due to the presence of synergic action of both major and minor bioactive components that inhibited hydrolytic enzymes (proteases) or suppressed partners cell wall envelop proteins, microbial adhesions, and non-specific interactions with carbohydrates (36). Further, antimicrobial potential of CFME against G- bacteria made an important discovery, as g-bacteria are mostly resistant to antibiotics. According to earlier research, G-bacteria have stronger cell walls than g+ bacteria, which enable them to show resistance to antibiotics. However, the absence of minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC) data represents a limitation of the present study, and inclusion of these parameters in future work would provide a more quantitative assessment of antibacterial efficacy.
Membrane Integrity Assay (Cytoplasmic Content Release Ccr) of C. Seifrizii Methanol Extracts
As postulated earlier, molecular docking predicted binding affinity of phytocompounds with cell wall based PdeF receptor and identifies key interactions (like hydrogen bonding) between a ligand and a target or receptor. To support the postulated mechanism of PdeF inhibition and the antibacterial action of CFME, a cell wall integrity test is often undertaken to illustrate how the drug affects bacterial structural defenses. It verifies if the antibacterial drug works by increasing membrane permeability or causing damage to the bacterial cell wall, rather than just metabolic inhibition. It aids in determining whether the extract's constituents target structural integrity or outer membrane proteins. Thereby to support the proposed mechanism of PdeF inhibition and antibacterial activity of CFME, cell wall integrity assay was conducted. The membrane permeability experiment was used to confirm the pathogen's cytoplasmic leakage upon engagement with the CFME formulations. Spectroscopy using UV light was used to measure the fluid system's absorption of visible and ultraviolet wavelengths. This helped to establish a connection between the amount of cytoplasmic leakage and light absorption. P. aeruginosa (MTCC 424), S. aureus (MTCC 3160), B. subtilis (MTCC 121), and E. coli (MTCC 40) were exposed to CFME for 0–12 h in order to examine the membrane integrity of these substances Supplementary Figure 4. Antibacterial action was demonstrated by CFME in a test pathogen-dependent manner. Additionally, all bacterial cells treated with CFME showed a linear rise in CCR as evidenced by increase in absorbance values. The results showed proving that the permeability of the bacterial cell wall was increased after 12 h of treatment with CFME. The leakage of nucleic acids or proteins from the cell is often a critical sign of cell membrane integrity. CFME showed almost greater CCR to reference control Triton-X-100 with MTCC 40, 121, and 3160 pathogens as compared to positive control data. By serving as an in vitro validation technique to demonstrate that the anticipated, in silico binding (docking) of a chemical to a target protein truly causes a physical, functional, or structural change in the membrane, membrane integrity assays validated the docking of bioactive substances. This prediction was functionally validated by the membrane integrity experiment. A ligand binding to a membrane-related enzyme may have been demonstrated by docking. The functional consequence of the binding, which is that the substance affects the structural integrity of the membrane, was validated by an experiment that measured the leakage of cytoplasmic components. Numerous new drugs and natural extracts have antibacterial activity through morphological alterations and membrane disruption. The bacterial membrane's integrity is essential for life and can either directly or indirectly causes cell death. The bacterial cell membrane performs important functions such as lipid production, transport, osmoregulation, and peptidoglycan cross-linking. Studies in the literature on the disruption of membrane cells by natural extracts from various medicinal plants have shown that natural extracts and their constituents act against a variety of targets, especially the cytoplasm and cell membrane, and sometimes completely change the Ammendolia morphology of the cells (37). In essence, while docking predicted that a compound binds to a site that might lead to disruption, the membrane integrity assay proved that the predicted interaction leads to actual membrane disruption. The assay provides physical evidence that CFME causes structural failure in the bacteria, validating that PdeF inhibition leads to, or coincides with, cell wall collapse.
In silico Pharmacokinetics
Supplementary Table 3, describes physiochemical properties and drug likeness of osthole. The Molecular Weight of osthole was 244.29 g/mol and topological Surface Area (TPSA) was 105.693 Ų. LogP value was 3.34. Drug likeness was evaluated by based on rules such as using Lipinski, Veber, and Egan's. It was observed that bioactive compound osthole did not violate Lipinski rule of five (RO5). Drugs used orally are not allowed to break more than one of the five Lipinski guidelines. Bioavailability score of osthole was 0.55. The pace and degree to which a medication (or other chemical) enters the bloodstream and circulates throughout the body is referred to as its bioavailability. It's a key component in figuring out how well a medication will function after being taken. A "good" bioavailability score, especially for oral drugs, generally indicates a high probability of a compound being absorbed into the bloodstream after being administered; a score of 0.55 or higher is often considered optimal, meaning a large portion of the drug will be absorbed and enter the systemic circulation (38). However, the ideal score can vary depending on the specific drug and its intended use. Thus, it can be said that the identified compound is soluble and permeable, making them suitable for usage as oral anticancer medications. These observations could be a good indicator of medication safety (39). This proved that molecule possessed properties similar to those of drugs.
Further ADMET properties of osthole were also predicted using pkCSM online server Supplementary Table 4. Pharmacokinetic properties include absorption (human intestinal absorption (HIA) and human colon adenocarcinoma cells (Caco-2), distribution (plasma protein binding (PPB) and blood-brain barrier (BBB)), metabolism (CYP2D6 and CYP3A4 as substrate and inhibition), and toxicity test (mutagenic and carcinogenic) were predicted in compounds of papaya leaves before further stages (clinical trials). Caco-2 showed value of -4.612, indicating good permeability. Caco-2 is widely used as a model in in vitro testing when predicting human drug absorption through the intestinal epithelial cell barrier. Osthole showed high GI absorption with TPSA 39.44 Å. Srimai et al. (2013) found that ligands with smaller molecular weights had a higher probability of diffusing rapidly and overcoming biological membranes. According to Wu et al. (39), TPSA is a very accurate indicator of drug absorption. A log BB value of 0.426 indicates a compound with high blood-brain barrier (BBB) permeability. BBB is used to characterize the distribution of compounds to the blood-brain barrier membrane permeability (41). A log Ps (logarithm of the permeability-surface area product) value of -2.01 indicates moderate to high, likely sufficient, CNS permeability. Further, osthole was non-substrate to P-glycoprotein (P-gp) efflux exporter. By pumping drugs back into the lumen, P-gp decreases the absorption of drugs in the stomach. Osthole had an inhibitory effect on the CYP1A2, CYP2C19 and CYP2C9 enzymes and non-inhibitory capabilities against the other enzyme series such as CYP2D6 and CYP3A4. According to Srimai et al. (40), the liver is home to the CYP450 class of enzymes that aid in the organism's detoxification process. CYP2D6 and CYP3A4 are two of the subtypes of cytochrome P450 (CYP450), an essential enzyme system for drug metabolism in the liver. Since they metabolize around 50% and 30% of the medications on the market, respectively, CYP3A4 and CYP2D6 are the most important CYP enzymes (41). Osthole was non-substrate to Renal OCT2 (Organic Cation Transporter 2). These are transporters, playing key roles in drug elimination. Radar and network charts to depict ADMET is also illustrated in Supplementary Figure 5.
Ligands' toxicity profile was also determined in Supplementary Table 4. According to Van Norman (42), a toxicity study is required for drugs that might endanger humans, animals, or the environment. Predication was based on different level of toxicity such as organ toxicity (neurotoxicity, cardiotoxicity, hepatotoxicity, respiratory toxicity, and nephrotoxicity), toxicological endpoints (like cytotoxicity, immunotoxicity, mutagenicity, carcinogenicity, clinical toxicity, ecotoxicity, and nutritional toxicity), toxicological pathways (AOPs), toxicity targets, targets for molecular initiating events (MIEs), and six molecular targets for metabolism are among the diverse levels of toxicity that the prediction scheme is divided into, thereby providing insights into the possible molecular mechanism behind such toxic response. Further, in-depth toxicological evaluation revealed that both of the bioactive substances used as ligand is insensitive to biological processes that depend on targets, including the stress response or nuclear receptor signaling pathways. These have been described as significant biological targets in the human body. It was observed that osthole is safe bioactive components since they fall within toxicity class 5, gives strong indications that this compound may nonetheless be non-toxic. The lethal toxicity dosage of a medicine is determined by the hazardous unit called LD50. Osthole expected lethal dose (LD50) was 2905mg/kg making it favorably less toxic. The higher the lethal dose per kg of a compound, the lesser its toxicity. The log10 (LD50) values of 3.6 for osthole on the hazardous scale demonstrated the bioactive chemical's non-toxic nature. The −log10 (LD50) value of ligands on the hazardous scale falls between 2.8 and 3.7, indicating that the bioactive compounds are not poisonous. The findings of the Ames test, a common biological test that uses mutant bacteria (Salmonella typhimurium/E. coli) to rapidly screen compounds for their ability to induce DNA mutations (mutagenicity) and, therefore, cancer (carcinogenicity), are referred to as Ames toxicity. Notably, Ames test for osthole was negative, suggesting that osthole was neither mutagen or carcinogenic. Tiwari et al. (43) state that if all medications are mutagenic, meaning they damage live cells and are the main cause of diseases like cancer. According to the toxicity profiles, which include hepatotoxicity, and skin Sensitisation, osthole was not harmful to any organs Supplementary Table 4.
The goal of Tox21 (Toxicology in the 21st Century) is to determine how environmental toxins disrupt human biological processes and employ high-throughput screening to link these disruptions to unfavorable health effects. In an effort to replace animal testing with molecular and computational data, it assesses substances against important pathways, such as nuclear receptors, stress response pathways, and developmental pathways. Interestingly, osthole was inactive against all pathways and receptors involved in different pathways Supplementary Table 4. Over all, it was observed that compounds were inactive for majority of pathways thus exerting non toxic effects.
Conclusion
This study reports phytochemistry and antibacterial potential of C. seifrizii fruit based methanol extracts. Chemical profiling revealed the presence of the major bioactive compound osthole (12.28%) in C. seifrizii fruit-based methanol extracts. In silico study revealed robust docking of major phytocompound osthole with PdeF proteins which was validated by MDS. In vitro antibacterial activity assays of C. seifrizii extracts demonstrated potential growth inhibition against tested pathogens. Therefore, we suggest that C. seifrizii extracts has potential to mitigate infections caused by g positive and g negative bacteria. To make space for these compounds in drug development, more research utilizing in vitro and in vivo models should be carried out to validate these compounds C. seifrizii fruit based methanol extracts.
Declarations
Acknowledgment
Authors acknowledge equipment lab established under FIST grant (level 0) supported by Dept. of Science and Technology (DST), Ministry of Science and Technology, Govt. of India
Conflict of Interest
The authors declare no conflicting interest.
Data Availability
The unpublished data is available upon request to the corresponding author.
Ethics Statement
Ethical approval was not required for this study.
Funding Information
This work was funded by the Department of Science and Technology, Government of India, DST/SEED/SCSP/STI/2019/253
Supplemental Material
Supplementary Tables and Figures are available and can be downloaded the <a class="cursor-pointer" href="https://etflin.com/file/document/20260407113058_485211_445c9e02.docx">following link</a>.
References
- Harris AD, Samore MH, Lipsitch M, Kaye KS, Perencevich E, Carmeli Y. Control‐Group Selection Importance in Studies of Antimicrobial Resistance: Examples Applied toPseudomonas aeruginosa,Enterococci, andEscherichia coli. Clin Infect Dis. 2002;34(12):1558-1563. doi: https://doi.org/10.1086/340533
- Garde S, Selvaraj H, Chandramouli A, Reddy GS, Bahety D, Chodisetti PK, et al. A conserved editing mechanism for the fidelity of bacterial cell wall biosynthesis. Proc. Natl. Acad. Sci. U.S.A. 2025;122(28)[No Pages]. doi: https://doi.org/10.1073/pnas.2505676122
- Lee MS, Hsieh KY, Kuo CI, Lee SH, Garde S, Reddy M, et al. Structural Basis for the Peptidoglycan-Editing Activity of YfiH. mBio. 2022;13(1)[No Pages]. doi: https://doi.org/10.1128/mbio.03646-21
- Agostini-Costa TDS. Bioactive compounds and health benefits of some palm species traditionally used in Africa and the Americas – A review. Journal of Ethnopharmacology. 2018;224[No Issue]:202-229. doi: https://doi.org/10.1016/j.jep.2018.05.035
- Rincón-Cervera MÁ, Lahlou A, Chileh-Chelh T, Lyashenko S, López-Ruiz R, Guil-Guerrero JL. Arecaceae Seeds Constitute a Healthy Source of Fatty Acids and Phenolic Compounds. Plants. 2023;12(2):226. doi: https://doi.org/10.3390/plants12020226
- Bernal R, Galeano G, García N, Olivares IL, Cocomá C. Uses and commercial prospects for the wine palm, Attalea butyracea, in Colombia. Ethnobot. Res. App. 2010;8[No Issue]:255. doi: https://doi.org/10.17348/era.8.0.255-268
- Inbathamizh L. Indoor medicinal plants: beneficial biocatalysts for air filtration and bioremediation—a review. Int J Green Pharm. 2020;14.
- Mohamed F, Ahmed H, El-A RA, Zeid AH. Chemopreventive activity of Chamaedorea sefrizii and Attalea butyracea leaf extracts in relation to its metabolite fingerprints analysed via UPLC-qTOF-MS and chemometrics. Biosci Res. 2019;16(4):3330–3338.
- Amrita, Kaur I, Sharma AD. Underutilized Plant Cymbopogan martinii Derived Essential Oil Is Excellent Source of Bioactives with Diverse Biological Activities. Russ. Agricult. Sci. 2023;49(1):100-117. doi: https://doi.org/10.3103/s1068367423010044
- Adams RP. Identification of essential oil components by gas chromatography/mass spectrometry. 4th ed. Carol Stream (IL): Allured Publishing; 2012.
- M S, P M. Phytochemical and UV spectrum Analysis of Azadirachta indica, Calotropis gigantea, and Ricinus communis. Pharma Innovation. 2023;12(6):2501-2504. doi: https://doi.org/10.22271/tpi.2023.v12.i6ac.20771
- Mabasa XE, Mathomu LM, Madala NE, Musie EM, Sigidi MT. Molecular Spectroscopic (FTIR and UV-Vis) and Hyphenated Chromatographic (UHPLC-qTOF-MS) Analysis and In Vitro Bioactivities of the Momordica balsamina Leaf Extract. Biochemistry Research International. 2021;2021[No Issue]:1-12. doi: https://doi.org/10.1155/2021/2854217
- Akiyama R, Suzuki K, Llave Y, Matsumoto T. Fluorescence Spectroscopy and a Convolutional Neural Network for High-Accuracy Japanese Green Tea Origin Identification. AgriEngineering. 2025;7(4):95. doi: https://doi.org/10.3390/agriengineering7040095
- Szukay B, Gałęcki K, Kowalska-Baron A, Budzyński J, Fisz JJ. Application of Steady-State and Time-Resolved Fluorescence Spectroscopy in Identification of Bee Products. Food Anal. Methods. 2024;17(9):1312-1326. doi: https://doi.org/10.1007/s12161-024-02667-w
- Donaldson L. Autofluorescence in Plants. Molecules. 2020;25(10):2393. doi: https://doi.org/10.3390/molecules25102393
- Singh V, Mishra AK. White Light Emission from Vegetable Extracts. Sci Rep. 2015;5(1). doi: https://doi.org/10.1038/srep11118
- Shukla U. Fourier transform infrared spectroscopy: A power full method for creating fingerprint of molecules of nanomaterials. Journal of Molecular Structure. 2025;1322:140454. doi: https://doi.org/10.1016/j.molstruc.2024.140454
- Mardare (Balusescu) G, Lazar L, Malutan T. Spectroscopic investigation and chemical fingerprint of Datura innoxia dry biomass. Comptes Rendus. Chimie. 2022;25(S3):227-235. doi: https://doi.org/10.5802/crchim.169
- Bouzidi A, Azizi A, Messaoudi O, Abderrezzak K, Vidari G, Hellal AN, et al. Phytochemical analysis, biological activities of methanolic extracts and an isolated flavonoid from Tunisian Limoniastrum monopetalum (L.) Boiss: an in vitro and in silico investigations. Sci Rep. 2023;13(1). doi: https://doi.org/10.1038/s41598-023-46457-6
- Duru C. Mineral and phytochemical evaluation of Zea mays husk. Scientific African. 2020;7:e00224. doi: https://doi.org/10.1016/j.sciaf.2019.e00224
- Yang Y, Song ZG, Liu ZQ. Synthesis and antioxidant capacities of hydroxyl derivatives of cinnamoylphenethylamine in protecting DNA and scavenging radicals. Free Radical Research. 2010;45(4):445-453. doi: https://doi.org/10.3109/10715762.2010.540576
- Zhang Z, Li X, Sang S, McClements DJ, Chen L, Long J, et al. Polyphenols as Plant-Based Nutraceuticals: Health Effects, Encapsulation, Nano-Delivery, and Application. Foods. 2022;11(15):2189. doi: https://doi.org/10.3390/foods11152189
- Chen L, Jin Y, Chen H, Sun C, Fu W, Zheng L, et al. Discovery of caffeic acid phenethyl ester derivatives as novel myeloid differentiation protein 2 inhibitors for treatment of acute lung injury. European Journal of Medicinal Chemistry. 2018;143:361-375. doi: https://doi.org/10.1016/j.ejmech.2017.11.066
- Agu PC, Afiukwa CA, Orji OU, Ezeh EM, Ofoke IH, Ogbu CO, et al. Molecular docking as a tool for the discovery of molecular targets of nutraceuticals in diseases management. Sci Rep. 2023;13(1). doi: https://doi.org/10.1038/s41598-023-40160-2
- Sahoo RN, Pattanaik S, Pattnaik G, Mallick S, Mohapatra R. Review on the use of Molecular Docking as the First Line Tool in Drug Discovery and Development. Ijps. 2022;84(5). doi: https://doi.org/10.36468/pharmaceutical-sciences.1031
- Meng XY, Zhang HX, Mezei M, Cui M. Molecular Docking: A Powerful Approach for Structure-Based Drug Discovery. Cad. 2011;7(2):146-157. doi: https://doi.org/10.2174/157340911795677602
- Pinzi L, Rastelli G. Molecular Docking: Shifting Paradigms in Drug Discovery. Ijms. 2019;20(18):4331. doi: https://doi.org/10.3390/ijms20184331
- Riu F, Ruda A, Ibba R, Sestito S, Lupinu I, Piras S, et al. Antibiotics and Carbohydrate-Containing Drugs Targeting Bacterial Cell Envelopes: An Overview. Pharmaceuticals. 2022;15(8):942. doi: https://doi.org/10.3390/ph15080942
- Zhydzetski A, Głowacka-Grzyb Z, Bukowski M, Żądło T, Bonar E, Władyka B. Agents Targeting the Bacterial Cell Wall as Tools to Combat Gram-Positive Pathogens. Molecules. 2024;29(17):4065. doi: https://doi.org/10.3390/molecules29174065
- Liu K, Watanabe E, Kokubo H. Exploring the stability of ligand binding modes to proteins by molecular dynamics simulations. J Comput Aided Mol Des. 2017;31(2):201-211. doi: https://doi.org/10.1007/s10822-016-0005-2
- Ishak S, Aris S, Halim K, Ali M, Leow T, Kamarudin N, et al. Molecular Dynamic Simulation of Space and Earth-Grown Crystal Structures of Thermostable T1 Lipase Geobacillus zalihae Revealed a Better Structure. Molecules. 2017;22(10):1574. doi: https://doi.org/10.3390/molecules22101574
- Elaissi A, Rouis Z, Salem NAB, Mabrouk S, ben Salem Y, Salah KBH, et al. Chemical composition of 8 eucalyptus species' essential oils and the evaluation of their antibacterial, antifungal and antiviral activities. BMC Complement Altern Med. 2012;12(1). doi: https://doi.org/10.1186/1472-6882-12-81
- Ammendolia DA, Bement WM, Brumell JH. Plasma membrane integrity: implications for health and disease. BMC Biol. 2021;19(1). doi: https://doi.org/10.1186/s12915-021-00972-y
- Cirillo N. A Roadmap for the Rational Use of Biomarkers in Oral Disease Screening. Biomolecules. 2024;14(7):787. doi: https://doi.org/10.3390/biom14070787
- Wu C, Liu Y, Yang Y, Zhang P, Zhong W, Wang Y, et al. Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods. Acta Pharmaceutica Sinica B. 2020;10(5):766-788. doi: https://doi.org/10.1016/j.apsb.2020.02.008
- Srimai V, Ramesh M, Satya Parameshwar K, Parthasarathy T. Computer-aided design of selective Cytochrome P450 inhibitors and docking studies of alkyl resorcinol derivatives. Med Chem Res. 2013;22(11):5314-5323. doi: https://doi.org/10.1007/s00044-013-0532-5
- Saptarini NM, Kelutur FJ, Corpuz MJAT. Exploring the Antidiarrheal Properties of Papaya Leaf: Insights In Vivo Study in Mice‐Model and In Silico Analysis at M3 Muscarinic Acetylcholine Receptor Interaction. Scientifica. 2024;2024(1). doi: https://doi.org/10.1155/2024/1558620
- Van Norman GA. Limitations of Animal Studies for Predicting Toxicity in Clinical Trials. JACC: Basic to Translational Science. 2020;5(4):387-397. doi: https://doi.org/10.1016/j.jacbts.2020.03.010
- Tiwari V, Shandily S, Albert J, Mishra V, Dikkatwar M, Singh R, et al. Corrigendum to “Insights into medication-induced liver injury: Understanding and management strategies” [Toxicol. Rep. 14 (2025) 101976]. Toxicology Reports. 2025;14:102068. doi: https://doi.org/10.1016/j.toxrep.2025.102068
- Nirmala MJ, Durai L, Rao KA, Nagarajan R. <p>Ultrasonic Nanoemulsification of <em>Cuminum cyminum</em> Essential Oil and Its Applications in Medicine</p>. Ijn. 2020;Volume 15:795-807. doi: https://doi.org/10.2147/ijn.s230893
- Lima SL, Colombo AL, de Almeida Junior JN. Fungal Cell Wall: Emerging Antifungals and Drug Resistance. Front. Microbiol. 2019;10. doi: https://doi.org/10.3389/fmicb.2019.02573