REVIEW
Genetic Polymorphism of N-acetyltransferase 2 (NAT2) among Patients with Tuberculosis: A Scoping Review of the Indonesian Studies
Academic Editor: Indriyati Hadi Sulistyaningrum
Sciences of Pharmacy|Vol. 5, Issue 2, pp. 129-137 (2026)
Received
Feb 16, 2026Revised
Mar 15, 2026Accepted
Mar 27, 2026Published
Apr 21, 2026
Abstract
Introduction
Tuberculosis (TB) is still an epidemic disease in the world, with the number of new cases increasing every year. According to the World Health Organization (WHO) report in 2024, more than 8.2 million people in the world were diagnosed with new TB cases in 2023, with 1.25 million deaths. The top five countries accounting for more than 59% of TB cases worldwide are India (26%), Indonesia (10%), China (6.8%), Philippines (6.8%), and Pakistan (6.3%) (1). The estimated incidence of TB in Indonesia in 2021 was 969, 000 or 354 per 100, 000, with a treatment success rate of 86.5% in 2022, and has not yet met the national target of 90%. One of the efforts made by the Indonesian Ministry of Health to support the elimination of TB in Indonesia by 2030 is to increase the treatment success rate to ≥ 90% (2). Treatment failure contributes to the occurrence of drug-resistant TB (DR-TB) cases, which are increasing every year in Indonesia. Therefore, optimizing antimicrobial therapy to reduce antituberculosis drug (ATD) resistance is crucial.
Isoniazid is one of the first-line ATDs to treat TB. It shows an early bactericidal effect by inhibiting mycolic acid synthesis, an important component of the TB bacterial cell wall. (3) According to the national tuberculosis guidelines, isoniazid is prescribed at 4-6 mg/kgBW per day and a maximum daily dose of 300 mg (4). However, isoniazid exhibits wide interindividual variability, which leads to alterations in its pharmacokinetic profile. A substantial proportion of patients with TB receiving standard dosing of Isoniazid have plasma drug concentrations below the predicted therapeutic range. For example, approximately 64–88% of patients were reported to have subtherapeutic isoniazid exposure in studies that measured plasma concentrations at 2 hours post-dose or evaluated pharmacokinetic parameters such as peak concentration (Cmax) and the area under the concentration–time curve (AUC). These findings highlight considerable interindividual variability in isoniazid pharmacokinetics among tuberculosis patients in Indonesia (5).
The antimycobacterial activity of isoniazid correlates with its plasma concentration. However, wide interindividual variability in isoniazid pharmacokinetics may lead to subtherapeutic drug concentrations, resulting in treatment failure and an increased risk of antibiotic resistance. (6) Subtherapeutic concentrations of isoniazid may result from both genetic and non-genetic factors. Non-genetic factors include poor treatment adherence, previous TB treatment (retreatment), inadequate drug dosing, and low body weight (7). Genetic factors contributing to interindividual variability in isoniazid pharmacokinetics primarily involve polymorphisms of the N-acetyltransferase 2 (NAT2) gene (6). Isoniazid undergoes hepatic metabolism mediated by the enzyme arylamine N-acetyltransferase 2 (NAT2) through an acetylation pathway. However, genetic polymorphisms in NAT2 lead to substantial interindividual variability in isoniazid metabolism and they were associated with treatment failure and drug resistance (8). Treatment failure and drug resistance are primarily associated with microbiological and treatment-related factors, such as inadequate drug exposure, poor adherence, or the presence of resistant strains of Mycobacterium tuberculosis. In contrast, hepatotoxicity represents an adverse drug reaction that is largely related to host factors, including genetic variability in drug metabolism (6, 7).
The distribution of NAT2 genetic polymorphisms varies considerably across ethnic groups worldwide, including in Indonesia. The distribution of NAT2 polymorphisms differs significantly across ethnic groups and geographic regions, as reported in previous pharmacogenetic studies, and this variation may influence the prevalence of slow, intermediate, and fast acetylator phenotypes. Such variability may affect the metabolism of isoniazid and consequently lead to differences in drug exposure and treatment-related outcomes among patients with TB (9-11). The severity of drug-induced liver injury (DILI) due to polymorphism of NAT2 was also different among studies (11, 12). Studies reported the prevalence of ATD-induced DILI ranged from 6.5% to 50% (13, 14). The impact of NAT2 genetic polymorphisms requires further investigation, particularly regarding their relationship with ethnicity and disease-specific differences, as well as their implications for individualized isoniazid dosing applicable in clinical practice among TB patients in Indonesia. Therefore, maintaining plasma isoniazid concentrations within the therapeutic range is essential to optimize treatment outcomes and minimize adverse effects.
There are differences among studies regarding the consistency and strength of the association between NAT2 polymorphisms and INH-DILI, despite growing evidence in Indonesia. These discrepancies may be attributed to differences in population genetics, diagnostic criteria, and study design. Thus, a comprehensive review is necessary to address the gap. A scoping review is considered the most appropriate approach to examine NAT2 genetic polymorphisms among patients with TB in Indonesia, as it enables a comprehensive mapping of existing evidence, including study designs, population characteristics, genotyping methods, and reported clinical outcomes. Furthermore, this approach facilitates current research gaps and provides a valuable foundation for future pharmacogenetic research on tuberculosis in the Indonesian population. Although the implementation of pharmacogenetics in the treatment of TB in Indonesia faces several challenges, including limitations in laboratory facilities, cost constraints, and healthcare infrastructure, advances in molecular technologies and the growing interest in precision medicine provide opportunities for broader integration of this approach in the future to improve the safety and effectiveness of TB therapy. Therefore, the objective of this scoping review is to map the existing evidence on NAT2 genetic polymorphisms among patients with tuberculosis in Indonesia.
Methodology
Search strategy and study selection
This scoping review was reported in accordance with the PRISMA Extension for Scoping Reviews (PRISMA-ScR) guidelines to ensure transparency, completeness, and reproducibility of the literature review process. These guidelines extend the Preferred Reporting Items for Systematic Reviews and Meta-Analyses and were specifically developed for scoping reviews (15). The search applies a combination of the following keyword variants: “tuberculosis”, “TB”, “isoniazid”, “INH”, “hepatotoxicity”, “drug-induced liver injury”, “polymorphism”, ”pharmacogenomic", “treatment outcomes”, “pharmacokinetic”, “N-acetyl-transferase 2”, “NAT2”, and “Indonesia”. The study includes literature indexed by Google Scholar, PubMed, and Scopus databases. Only original, peer-reviewed articles published in English or Indonesian and conducted in Indonesia between 2011 and 2024 were included in this study. These terms were combined using Boolean operators with "OR", "AND", and "NOT" to retrieve articles more specifically.
The search in the PubMed databases was conducted using the following combination of terms ("N-acetyltransferase 2" OR NAT2 OR "acetylator status" OR "NAT2 polymorphism") AND (tuberculosis OR TB OR "Mycobacterium tuberculosis") AND ("Drug-Induced Liver Injury" OR hepatotoxicity OR "liver toxicity") AND (Indonesia OR Indonesian). The search in the Scopus and Google Scholar databases was conducted using the following combination of terms ("NAT2" OR "N-acetyltransferase 2" OR "NAT2 polymorphism" OR "acetylator status" OR "slow acetylator" OR "fast acetylator") AND ("tuberculosis" OR "TB" OR "Mycobacterium tuberculosis") AND ("hepatotoxicity" OR "drug-induced liver injury" OR DILI OR "liver injury" OR "liver toxicity")
Inclusion and exclusion criteria
The inclusion criteria in this scoping review were: 1) articles written either in English or Indonesian; 2) articles as original articles with cross-sectional, cohort, or case-control studies, and pharmacokinetic studies; 3) reporting genetic polymorphism of NAT2 and or NAT2 phenotypic (acetylator status). Articles analyzing the association between the genetic polymorphism of NAT2 with drug-induced liver injury, pharmacokinetic profile, and treatment outcomes were also included. Studies that included patients receiving INH for conditions other than tuberculosis, such as other mycobacterial infections, were excluded. Reviews, letters to the editor, and commentaries were also excluded from this review.
Ethical Statement
The current study did not require ethical approval because it was a scoping review; therefore, informed consent was waived.
Process of study selection
The article selection process was conducted systematically using the web-based application Rayyan, which is designed to facilitate the literature screening process in both systematic and scoping reviews. All references retrieved from various electronic databases were first imported into reference management software to identify and remove duplicate records and were subsequently exported to Rayyan for further screening. The study selection process was conducted in two stages. The first stage involved screening titles and abstracts to assess eligibility based on the predefined inclusion criteria. The second stage consisted of a full-text review of articles that passed the initial screening. The entire selection process was conducted independently by two researchers (ONP, SRR), and any disagreements were resolved through discussion or, when necessary, by consulting a third researcher (YY, JJ, AYN). Articles considered relevant or potentially relevant were then advanced to the full-text review stage to determine their final eligibility for inclusion in the review. The study selection process was reported using the PRISMA-ScR flow diagram.Data extraction
Data from studies that met the inclusion criteria were extracted using a standardized data extraction form. The form was designed to collect relevant information from each included article, including general study characteristics, research methods, population characteristics, and key findings related to the objectives of the scoping review. The author’s name, study location, year of publication, sample size, study design, and summary of findings were extracted to provide a descriptive overview of each study. The extracted data also included the acetylator phenotype and the frequency of single-nucleotide polymorphism (SNP) alleles. Because the study's findings were diverse, they were presented as a narrative synthesis. Aspartate aminotransferase (AST) or alanine aminotransferase (ALT) levels greater than five times the upper limit of normal (ULN), AST or ALT levels greater than three times the ULN with symptoms like nausea, vomiting, lack of appetite, abdominal discomfort, or jaundice, or AST or ALT levels greater than three times the ULN with total bilirubin levels greater than two times the ULN were the criteria for diagnosing DILI (16). Therefore, the standardized diagnostic criteria are necessary in the included studies to ensure consistency in the identification and reporting of hepatotoxicity cases associated with antituberculosis drugs. The outcomes were grouped into thematic categories, including genetic characteristics (NAT2 genotype and acetylator status), pharmacokinetic outcomes (isoniazid plasma concentration), and clinical outcomes (hepatotoxicity or treatment response), to facilitate a structured presentation of the evidence.
In this study, no quality assessment of the included studies was conducted because the primary objective of a scoping review is to map the available scientific evidence, describe study characteristics, and identify knowledge gaps rather than evaluate the effectiveness of interventions or produce quantitative effect estimates, as is typically done in a systematic review. In addition, the PRISMA-ScR reporting guidelines state that study quality assessments may be reported if conducted, but they are not required for a scoping review (15).
Results and Discussion
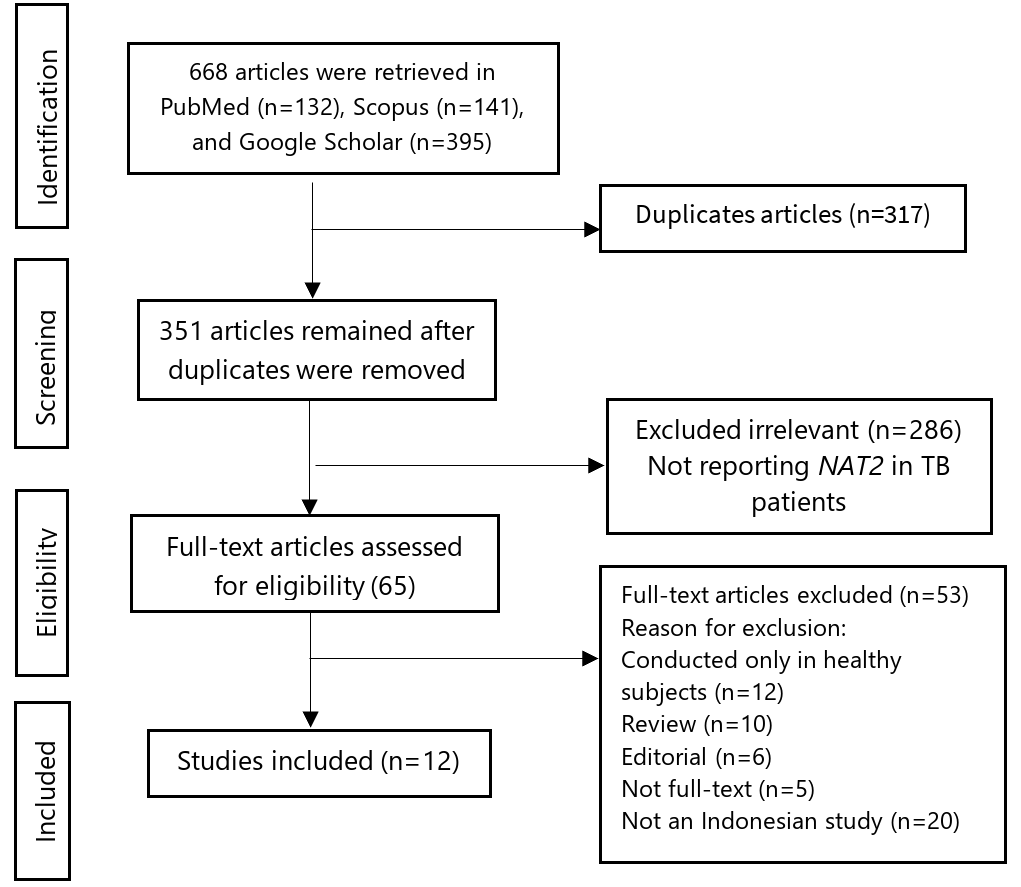
Using a combination of keywords, a total of 813 articles were identified across three databases. Of these, 317 duplicate records were removed. Of the remaining 496 articles, 286 were excluded after title and abstract screening, leaving 65 for full-text review. Subsequently, 50 articles were further excluded for various reasons. Finally, according to the PRISMA ScR flowchart, 12 articles were included in the scoping review. Figure 1 presents the PRISMA flowchart for article selection, while Supplemental Table 1 summarizes the characteristics of the studies included in this review.
We retrieved 12 articles for final analysis in this study. Our review found only one study reporting the NAT2 genetic polymorphism in pediatric TB. Three studies did not analyze the association between NAT2 genetic polymorphism and DILI or serum INH concentration (17, 18, 23). In this review, no studies involving TB patients were conducted in Kalimantan, Sulawesi, Bali, or Papua. Isoniazid is a prodrug that undergoes bioactivation by the KatG enzyme in Mycobacterium tuberculosis cells, forming an isoniazid–NAD (INH–NAD) adduct that competitively inhibits InhA, an enzyme involved in the fatty acid synthase (FAS) II pathway that is responsible for mycolic acid synthesis. This inhibition prevents mycolic acid production, ultimately leading to bacterial cell death (7). NAT2 is an enzyme encoded by the intronless NAT2 gene, which spans approximately 870 base pairs and is located on chromosome 8. Variations in the DNA sequence of the NAT2 gene, known as single-nucleotide polymorphisms (SNPs), result in differences in genotypes, haplotypes, and corresponding phenotypes. Based on their capacity to metabolize isoniazid, NAT2 haplotypes are phenotypically classified into fast, intermediate, and slow acetylators. Fast acetylators acetylate isoniazid five to six times faster than slow acetylators. The acetylation rate constant for monoacetylhydrazine is estimated to be fourfold greater in fast acetylators than in slow acetylators (25).
There are at least seven SNPs in the NAT2 gene, such as nucleotide change (nt) 191G>A at rs1801279, nt282C>T at rs1041983, nt 341T>C at rs1801280, nt 803A>G at rs1208, nt 590G>A at rs1799930, nt 857>A at rs1799931, and nt 481C>T at rs1799929. The distribution of these SNPs has been documented in international databases and reported to vary across different ethnic populations worldwide (26). The combination of multiple SNPs determines NAT2 haplotypes, leading to a wide range of acetylation phenotypes that influence NAT2 enzyme activity. For example, two NAT2*4 normal alleles or homozygous wild-type in the form of NAT*2/*4 are classified as fast acetylators. One NAT2*4 normal allele in combination with one NAT2*5, NAT-2*6, or NAT-2*7 mutant allele is classified as an intermediate acetylator. Two mutant alleles with genotype combinations such as nt481C> T, nt857G> A, or nt 590G> A in the form of NAT2*5, NAT2*6, or NAT2*7 are classified as slow acetylators (27).
We found one study by Pramono et al., who suggested that in TB patients with and without diabetes, variations in the NAT2 acetylator phenotype were present in both groups, with no evidence that diabetes alters acetylator status. TB patients with diabetes have a higher risk of DILI, but this is typically associated with metabolic and chronic inflammatory factors associated with diabetes, rather than changes in acetylator phenotype (17). However, the main limitations of the study include the very small sample size, limited geographic coverage, the absence of pharmacokinetic data or reported DILI events, and a cross-sectional design that restricts causal inference. Therefore, larger, multicenter studies incorporating clinical and pharmacokinetic data are warranted to generate more robust and generalizable evidence.
A study by Sahiratmaja et al., conducted in Kupang, West Nusa Tenggara, demonstrated that rapid and intermediate acetylators were more prevalent among TB patients (18, 23). A study by Perwitasari et al., conducted in Yogyakarta and Lampung, reported that more than 60% of TB patients were slow acetylators (9, 19). Acetylhydrazine (AcHz) is considered a key metabolite responsible for the hepatotoxic effects of isoniazid (28). Slow acetylators are at greater risk of accumulating toxic isoniazid metabolites, including AcHz, due to reduced acetylation capacity. In addition, unacetylated isoniazid may be hydrolyzed by amidase enzymes to form hydrazine, another hepatotoxic metabolite that contributes to liver injury (29).
Conversely, more than 60% of TB patients were rapid acetylators as reported by Anggraini et al. It was more likely because they used the sulphadimidine method to assess acetylation status (30). Meanwhile, most studies in this review used DNA sequencing to detect SNPs and the NAT2 phenotype. Different methods of assessing acetylator status may lead to different results. The study's small sample size, observational design, and lack of comprehensive pharmacokinetic and hepatotoxicity data limit its ability to provide robust insights into the impact of acetylation status on isoniazid serum concentrations and sputum conversion. These limitations underscore the need for larger, multicenter studies with more detailed genetic and clinical assessments to enhance the generalizability of findings and support causal inference.
Previous studies on several Indonesian ethnic groups in healthy subjects, including Javanese-Sundanese, Buginese, and Batak, also showed that a significant proportion of the population are slow acetylators (31-33). In contrast, a study by Sahiratmaja et al., reported that nearly 50% of household contacts of TB patients in Kupang, a region in eastern Indonesia, were rapid acetylators (18). It demonstrated substantial variations in NAT2 and acetylation status among Indonesia's ethnic groups. The distribution of NAT2 acetylator status varies across regions in Indonesia due to differences in the frequency of NAT2 alleles associated with ethnicity, influenced by population history, migration, and local genetic variation. This variation should be considered in individualized TB treatment strategies, including isoniazid dosing and monitoring for the risk of hepatotoxicity. Nonetheless, the frequency of inferred slow acetylators in the Indonesian population is relatively comparable to that observed in other Southeast Asian populations (34-36).
Seven studies reported an association between NAT2 genetic polymorphisms determining acetylator status and the occurrence of drug-induced liver injury (DILI) (9, 11, 12, 19-21, 24). TB patients with a slow acetylator phenotype were associated with DILI. Nevertheless, no studies report the onset of DILI in TB patients. The study by Perwitasari et al., found that 69.5%, 5.6%, and 25.0% of drug-sensitive pulmonary TB (DS-TB) patients were classified as slow, intermediate, and fast acetylators, respectively, and that the slow acetylator phenotype was significantly associated with DILI. TB patients with serum isoniazid concentrations ≥ 7 mcg/mL had a 1.2-fold higher risk of drug-induced liver injury (DILI) (9).
Another study by Yuliwulandari et al., conducted in DKI Jakarta, East Java, and Banten provinces, reported that TB patients with the slow acetylator phenotype had a 3.6-fold higher risk of developing drug-induced liver injury (DILI) compared with those with the fast acetylator phenotype (20). However, none of the studies analyzed several factors that may increase the risk of DILI, such as TB patients with HIV, smoking status, and history of alcohol drinking.
Only one study by Soedarsono et al., reported that TB patients with low BMI and slow acetylators were significantly associated with DILI (12). Patients with low BMI often have deficiencies in plasma proteins, including albumin, which play a critical role in the distribution of isoniazid, rifampicin, and their metabolites (37). Reduced plasma protein levels increase the free fraction of the drug, leading to greater hepatocyte exposure to toxic metabolites. Isoniazid and rifampicin are metabolized in the liver via acetylation and glucuronidation pathways. Consequently, malnutrition may alter hepatic enzyme activity and impair the detoxification of toxic metabolites such as hydrazine (38). This indicates that nutritional status is an important host factor that modulates the risk of hepatotoxicity associated with ATDs. Thus, further studies in Indonesia need to be conducted with attention to some covariates that may lead to an increased risk of DILI and alter the pharmacokinetic profile of isoniazid in TB patients.
The association between slow acetylator status and drug-induced liver injury (DILI) reported in Indonesian studies is supported by several studies conducted in India (39) and Thailand (35, 40), which demonstrated that slow acetylators have significantly higher serum isoniazid (INH) concentrations than fast and intermediate acetylators. Interestingly, a study in Thailand reported that among slow acetylators, individuals with the NAT2*6A/6A genotype had higher serum isoniazid concentrations than those with NAT2*6A/*6A, NAT2*5B/*6A, NAT2*5B/*5B, NAT2*6A/*7B, and NAT2*6A genotypes (35). This finding is consistent with our review, which demonstrated that the NAT26 polymorphism was more prevalent among TB patients with the slow acetylator phenotype.
Patients carrying the NAT2*6A allele are likely to have reduced NAT2 enzyme activity, which slows the metabolism of isoniazid and leads to the accumulation of toxic metabolites, thereby directly increasing the risk of DILI. Identifying patients with the NAT26A allele enables dose adjustment of isoniazid, closer monitoring of liver function throughout therapy, and a personalized (precision medicine) treatment approach, particularly in populations with a high prevalence of NAT26A. Therefore, NAT2 genotyping, specifically targeting the NAT2*6A allele, may serve as a safe and evidence-based strategy for preventing hepatotoxicity in Indonesia.
In our review, we identified an association between acetylation status and serum isoniazid (INH) concentration, as reported by Soedarsono et al. Rapid acetylators had significantly lower peak concentrations and area under the curve (AUC₀–₂₄) compared with patients with intermediate and slow acetylator phenotypes. However, the study was conducted in a single hospital in Surabaya, where the population is predominantly Javanese; therefore, ethnicity could not be included as a covariate (10). Consequently, different results may be observed when these findings are applied to other ethnic groups in Indonesia. Furthermore, the study did not include several covariates that may influence serum isoniazid concentration or clearance. Previous studies have reported that factors such as smoking status, previous exposure to first-line anti-TB treatment, elevated uric acid levels, age, and body mass index may be associated with serum INH concentrations (41-44). A study by Anggraini et al., demonstrated that serum isoniazid concentrations were approximately twice as high in slow acetylators as in fast acetylators. Nevertheless, acetylation status was not significantly associated with treatment outcomes, possibly due to the relatively small sample size, which limited the statistical analysis (30). In contrast, another study with a larger sample size reported that early treatment failure was more frequent in fast acetylators than in slow acetylators. Additionally, a study by Ulanova et al., found that isoniazid concentrations < 3 mcg/mL in rapid acetylators were associated with positive sputum cultures in the second month of treatment (45).
We identified two studies that suggested that MDR-TB patients were more likely to exhibit the rapid acetylator phenotype (21, 22). This suggests that the rapid acetylator phenotype may increase the likelihood of isoniazid resistance. Isoniazid is included in the shorter MDR-TB treatment regimen (9–11 months) at a daily dose of 450–600 mg. A study by Yunivita et al. involving Indonesian MDR-TB patients reported that approximately one-fourth of patients had AUC₀–₂₄/MIC and Cmax/MIC values below the proposed therapeutic targets. The proportion of fast acetylators was higher than that of slow acetylators (46). Although several limitations should be considered, the studies by Yuliwulandari et al., and Senja et al., provide valuable insights into the role of NAT2 polymorphisms in MDR-TB patients. As both studies employed observational designs, establishing a causal relationship between NAT2 variants and clinical outcomes remains challenging. In addition, the relatively small sample sizes may reduce statistical power and limit the generalizability of the findings. The pharmacokinetic evaluation was also limited, as the area under the curve, drug clearance, and half-life were not thoroughly assessed. Furthermore, important clinical outcomes, including hepatotoxicity and treatment response, were not consistently evaluated, and potential confounding factors such as nutritional status, comorbidities, and treatment adherence were not comprehensively examined. Therefore, larger and multicenter studies involving diverse populations in Indonesia are needed to evaluate the distribution of NAT2 phenotypes and their associations with both MDR-TB treatment efficacy and the risk of hepatotoxicity.
Pharmacogenomic screening for NAT2 polymorphisms has been proposed as a potentially cost-effective strategy to optimize isoniazid dosing and prevent DILI. A modeling study reported that NAT2 genotype-guided therapy gained approximately 19–24 quality-adjusted life years (QALYs) per 1.000 patients. In several high TB-burden countries, the incremental cost-effectiveness ratios ranged from $476 to $1, 780 per QALY gained. As these values fall below the national gross domestic product (GDP) per capita thresholds, NAT2 genotyping would be considered highly cost-effective (47). Although comparable pharmacoeconomic analyses have not yet been conducted in Indonesia, the high frequency of NAT2 polymorphisms and the substantial burden of antituberculosis drug-induced hepatotoxicity suggest that pharmacogenomic screening may represent a promising strategy for early risk detection and personalized TB therapy.
This review has several important limitations that should be carefully considered. First, the number of studies examining the impact of NAT2 genetic polymorphisms on therapeutic outcomes in TB patients remains relatively limited. Consequently, the currently available evidence is still insufficient to determine the direct effect of these polymorphisms on treatment failure among TB patients in Indonesia. Second, studies investigating multiple clinical and genetic factors that may influence isoniazid pharmacokinetics and hepatotoxicity are also scarce and limited in scope.
Conclusions
This scoping review indicates that N-acetyltransferase 2 (NAT2) gene polymorphisms play a significant role in determining acetylator phenotypes, which influence isoniazid metabolism and clinical outcomes among tuberculosis patients in Indonesia. Most evidence suggests that the slow acetylator phenotype is relatively prevalent in the TB patient population and is consistently associated with higher serum isoniazid concentrations and an increased risk of drug-induced liver injury during antituberculosis therapy. Certain genetic variants, particularly the NAT2*6A allele, have been reported to reduce NAT2 enzyme activity, thereby slowing isoniazid metabolism and leading to the accumulation of hepatotoxic metabolites, which increases susceptibility to hepatotoxicity. Conversely, patients with the rapid acetylator phenotype tend to have lower isoniazid concentrations due to increased drug clearance, which may result in subtherapeutic drug exposure and potentially affect therapeutic response. However, the available evidence remains limited because many studies involve relatively small sample sizes, restricted geographic coverage, and a lack of integration of genetic, pharmacokinetic, and clinical outcome data. Therefore, large-scale, multicenter prospective studies involving diverse ethnic populations across Indonesia are needed to generate more comprehensive evidence on the relationship between NAT2 polymorphisms, isoniazid metabolism, and TB treatment outcomes. Additionally, pharmacoeconomic evaluations are necessary to assess the feasibility of implementing NAT2 genotype screening as part of a precision medicine–based approach to TB treatment. The integration of pharmacogenomic information into clinical practice has the potential to improve treatment safety, reduce the risk of hepatotoxicity, and optimize the effectiveness of TB therapy in Indonesia.
Abbreviations
SNP: single nucleotide polymorphism; DILI: drug-induced liver injury; TB: tuberculosis; MDR-TB: multidrug-resistant tuberculosis; OR; odds ratio; 95% CI: 95% of confidence interval; INH: isoniazid; RIF: rifampicin; PZA: pyrazinamide; EMT: ethambutol; kgBW: kilogram body weight; NA: not available; NAT2: N-acetyltransferase 2; RR: relative risk; AUC: area under curve; QALY: quality-adjusted life years; MIC: minimum inhibitory concentration; Cmax: maximum concentration; ATDs: antitubercular drugs.
Declarations
Conflict of Interest
The authors declare no conflicting interest.
Data Availability
The data generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Ethics Statement
Ethical approval was not required for this study.
Funding Information
The authors declare that no financial support was received for the research, authorship, and/or publication of this article.
Supplemental Material
The <a class="cursor-pointer" href="https://etflin.com/file/document/20260406003138_923265_10786c3d.docx">Supplementary Data</a> section contains <b>Supplemental Table 1</b>. Characteristics of studies included in this scoping review.
References
- World Health Organization. WHO Global Tuberculosis Report 2024. Geneva. Switzerland.
- Ministry of Health of the Indonesian Republic. Tuberculosis Control Program Report in 2022. Ministry of Health of the Indonesian Republic. Jakarta. 2023.
- Dooley KE, Miyahara S, von Groote-Bidlingmaier F, Sun X, Hafner R, Rosenkranz SL, et al. Early Bactericidal Activity of Different Isoniazid Doses for Drug-Resistant Tuberculosis (INHindsight): A Randomized, Open-Label Clinical Trial. American Journal of Respiratory and Critical Care Medicine. 2020;201(11):1416-1424. doi: https://doi.org/10.1164/rccm.201910-1960oc
- Ministry of Health of the Indonesian Republic. National Guidelines for Tuberculosis Management. Ministry of Health of the Indonesian Republic. Jakarta. 2020
- Widhiartini IAA, Wirasuta MAG, Sukrama DM, Rai IBN. Therapeutic drug monitoring of rifampicin, isoniazid, and pyrazinamide in newly-diagnosed pulmonary tuberculosis outpatients in Denpasar area. Bali Med J. 2019;8(1):107. doi: https://doi.org/10.15562/bmj.v8i1.1304
- Shabani S, Farnia P, Ghanavi J, Velayati AA, Farnia P. Pharmacogenetic Study of Drugs Affecting Mycobacterium tuberculosis. The International Journal of Mycobacteriology. 2024;13(2):206-212. doi: https://doi.org/10.4103/ijmy.ijmy_106_24
- Erwin ER, Addison AP, John SF, Olaleye OA, Rosell RC. Pharmacokinetics of isoniazid: The good, the bad, and the alternatives. Tuberculosis. 2019;116:S66-S70. doi: https://doi.org/10.1016/j.tube.2019.04.012
- Mahajan R, Tyagi AK. Pharmacogenomic insights into tuberculosis treatment shows the NAT2 genetic variants linked to hepatotoxicity risk: a systematic review and meta-analysis. BMC Genom Data. 2024;25(1). doi: https://doi.org/10.1186/s12863-024-01286-y
- Wilffert B, Perwitasari D, Darmawan E, Mulyani U, Vlies PD, Alffenaar JW, et al. Polymorphisms of NAT2, CYP2E1, GST, and HLA related to drug-induced liver injury in indonesian tuberculosis patients. Int J Mycobacteriol. 2018;7(4):380. doi: https://doi.org/10.4103/ijmy.ijmy_143_18
- Soedarsono S, Jayanti RP, Mertaniasih NM, Kusmiati T, Permatasari A, Indrawanto DW, et al. Development of population pharmacokinetics model of isoniazid in Indonesian patients with tuberculosis. International Journal of Infectious Diseases. 2022;117:8-14. doi: https://doi.org/10.1016/j.ijid.2022.01.003
- Mohamed Noor NF, Salleh MZ, Mohd Zim MA, Bakar ZA, Fakhruzzaman Noorizhab MN, Zakaria NI, et al. NAT2 Polymorphism and Clinical Factors that Increase Antituberculosis Drug-Induced Hepatotoxicity. Pharmacogenomics. 2022;23(9):531-541. doi: https://doi.org/10.2217/pgs-2022-0022
- Hiranburana N, Sophonphan J, Kerr SJ, Ubolyam S, Usawakidwiree W, Torvorapanit P, et al. NAT 2 Polymorphisms and Antituberculosis-Induced Hepatotoxicity in Thai People Living With HIV: Insights From a Pharmacogenetic–Pharmacokinetic Cohort Study. The Journal of Infectious Diseases. 2025;233(2):287-297. doi: https://doi.org/10.1093/infdis/jiaf511
- Noviansyah EI, Djaharuddin I, Putrawan HA, Madolangan J, Natsir B, Pandu E. The Effect of Hepatoprotectors on the Risk of Drug-induced Hepatitis in Pulmonary Tuberculosis Patients. The International Journal of Mycobacteriology. 2025;14(3):219-225. doi: https://doi.org/10.4103/ijmy.ijmy_67_25
- Azmi RN, Ardiani T, Siami S. Differences in transaminase enzyme values and hepatoxic severity in antituberculosis drug use cross-sectional study in tuberculosis patients and tuberculosis-Human Immunodeficiency Virus co-infected. Med Hosp. 2022;9(1):37-42
- Supplemental Information 2: Preferred Reporting Items for Systematic reviews and Meta-Analyses extension for Scoping Reviews (PRISMA-ScR) Checklist. doi: https://doi.org/10.7717/peerj.10163/supp-2
- Bessone F, Hernandez N, Tagle M, Arrese M, Parana R, Méndez-Sánchez N, et al. Drug-induced liver injury: A management position paper from the Latin American Association for Study of the liver. Annals of Hepatology. 2021;24:100321. doi: https://doi.org/10.1016/j.aohep.2021.100321
- Pramono AA, Penggoam S, Sahiratmadja E, Utami NV, Achmad TH, Panigoro R. Status Asetilator Gen NAT2 pada Pasien Tuberkulosis dan Tuberkulosis dengan Diabetes Melitus di Kupang, Nusa Tenggara Timur. mkb. 2017;49(1):61-66. doi: https://doi.org/10.15395/mkb.v49n1.989
- Sahiratmadja E, Penggoam S, Maskoen AM, Pramono AA, Aryani D, Rahayu NS, et al. Distribution of rs1801279 and rs1799930 Polymorphisms in NAT2 Gene among Population in Kupang, Nusa Tenggara Timur, Indonesia. Indones Biomed J. 2018;10(1):56. doi: https://doi.org/10.18585/inabj.v10i1.330
- Perwitasari DA. Genotype polymorphisms of nat2 and cyp2e1 genes associated with drug induced liver injury (dili) in indonesian tuberculosis patients. Indonesian J. Pharm. 2016;27(1):22. doi: https://doi.org/10.14499/indonesianjpharm27iss1pp22
- Yuliwulandari R, Prayuni K, Susilowati RW, Subagyo, Soedarsono, M Sofro AS, et al. NAT2 Slow Acetylator is Associated with Anti-tuberculosis Drug-induced Liver injury Severity in Indonesian Population. Pharmacogenomics. 2019;20(18):1303-1311. doi: https://doi.org/10.2217/pgs-2019-0131
- Yuliwulandari R, Prayuni K, Razari I, Susilowati RW, Zulhamidah Y, Soedarsono S, et al. Genetic Characterization of N -acetyltransferase 2 variants in Acquired Multidrug-Resistant Tuberculosis in Indonesia. Pharmacogenomics. 2021;22(3):157-163. doi: https://doi.org/10.2217/pgs-2020-0163
- Senja M, Basyar M, Sabri YS, Afriani A. Correlation between N-Acetyltransferase 2 (NAT2) Polymorphism Genotype with Plasma Isoniazid (INH) Concentration in MDR TB Patients Receiving Short Regimen in West Sumatera. J Respirol Indones. 2022;42(1):26-33. doi: https://doi.org/10.36497/jri.v42i1.283
- Sahiratmadja E, Rini IA, Penggoam S, Charles A, Maskoen AM, Parwati I. Acetylator Status Among Newly Diagnosed and Recurrent Tuberculosis Patients from Kupang, Eastern Part of Indonesia. Pgpm. 2021;Volume 14:737-744. doi: https://doi.org/10.2147/pgpm.s311952
- Headriawan A, Pramono AA, Sukadi A, Chairulfatah A, Maskoen AM, Nataprawira HM. NAT2 Gene rs1041983 is Associated with Anti-Tuberculosis Drug Induced Hepatotoxicity Among Pediatric Tuberculosis in Bandung, Indonesia. Tacg. 2021;Volume 14:297-303. doi: https://doi.org/10.2147/tacg.s303668
- Imam F, Sharma M, Khayyam KU, Khan MR, Ali MD, Qamar W. Determination of isoniazid acetylation patterns in tuberculosis patients receiving DOT therapy under the Revised National tuberculosis Control Program (RNTCP) in India. Saudi Pharmaceutical Journal. 2020;28(6):641-647. doi: https://doi.org/10.1016/j.jsps.2020.04.003
- Cheng F, Jiang XG, Zheng SL, Wu T, Zhang Q, Ye XC, et al. N-acetyltransferase 2 genetic polymorphisms and anti-tuberculosis-drug-induced liver injury: a correlation study. Front. Pharmacol. 2023;14. doi: https://doi.org/10.3389/fphar.2023.1171353
- Lopes MQP, Teixeira RLF, Cabello PH, Nery JAC, Sales AM, Nahn J. R. EP, et al. Human N-acetyltransferase 2 (NAT2) gene variability in Brazilian populations from different geographical areas. Front. Pharmacol. 2023;14. doi: https://doi.org/10.3389/fphar.2023.1278720
- Ky Anh N, My Tung P, Kim MJ, Phuoc Long N, Cho YS, Kim DH, et al. Quantitative Analysis of Isoniazid and Its Four Primary Metabolites in Plasma of Tuberculosis Patients Using LC-MS/MS. Molecules. 2022;27(23):8607. doi: https://doi.org/10.3390/molecules27238607
- Sankar J, Chauhan A, Singh R, Mahajan D. Isoniazid-historical development, metabolism associated toxicity and a perspective on its pharmacological improvement. Front. Pharmacol. 2024;15. doi: https://doi.org/10.3389/fphar.2024.1441147
- Anggraini DI, Kristin E, Dwiprahasto I. The influence of acetylation status of tuberculosis patients on the isoniazid serum concentrations and sputum conversion after intensive phase therapy. JMedScie. 2018;50(01):59-69. doi: https://doi.org/10.19106/jmedsci005001201807
- Rahardjo DN, Mutiara YM, Wahjudi M, Andrajati R, Sartika RAD. The interaction of isoniazid on 3D structure variation of NAT2 gene and its effect on drug blood levels. J Med Chem Sci. 2024;7:1969-81
- Susilowati RW, Prayuni K, Razari I, Bahri S, Yuliwulandari R. High frequency of NAT2 slow acetylator alleles in the Malay population of Indonesia: an awareness to the anti-tuberculosis drug induced liver injury and cancer. Med J Indones. 2017;26(1):7-13. doi: https://doi.org/10.13181/mji.v26i1.1563
- Pasaribu E, Paranginangin JM, Dewi LVI. Identification of N-acetyltransferase 2 (NAT) gene polymorphisms in Batak. Health Inf J Penelit. 2023;15(Suppl 1):923-30
- Thaw KS, Choppradit C, Kijsanayotin P, Rodsiri R. Association of N-acetyltransferase 2 (NAT2) polymorphisms and elevated liver enzymes of Myanmar tuberculosis patients in Thailand. The Thai Journal of Pharmaceutical Sciences. 2020;44(4):210-215. doi: https://doi.org/10.56808/3027-7922.2455
- Ungcharoen U, Sriplung H, Mahasirimongkol S, Chusri S, Wichukchinda N, Mokmued P, et al. The Influence of NAT2 Genotypes on Isoniazid Plasma Concentration of Pulmonary Tuberculosis Patients in Southern Thailand. Tuberc Respir Dis. 2020;83(Supple 1):S55-S62. doi: https://doi.org/10.4046/trd.2020.0068
- Chan SL, Chua APG, Aminkeng F, Chee CBE, Jin S, Loh M, et al. Association and clinical utility of NAT2 in the prediction of isoniazid-induced liver injury in Singaporean patients. PLoS ONE. 2017;12(10):e0186200. doi: https://doi.org/10.1371/journal.pone.0186200
- Surarak T, Chumnumwat S, Nosoongnoen W, Tragulpiankit P. Efficacy, safety, and pharmacokinetics of isoniazid affected by NAT2 polymorphisms in patients with tuberculosis: A systematic review. Clinical Translational Sci. 2024;17(4). doi: https://doi.org/10.1111/cts.13795
- Liu Q, Huang L, Yan H, Zong Z, Chen Z, Wu X, et al. Clinical risk factors for moderate and severe antituberculosis drug-induced liver injury. Front. Pharmacol. 2024;15. doi: https://doi.org/10.3389/fphar.2024.1406454
- Munshi R, Panchal F, Desai U, Utpat K, Rajoria K. A study of N-acetyltransferase 2 gene polymorphisms in the Indian population and its relationship with serum isoniazid concentrations in a cohort of tuberculosis patients. Monaldi Arch Chest Dis. 2024;95(4). doi: https://doi.org/10.4081/monaldi.2024.3181
- Suvichapanich S, Wattanapokayakit S, Mushiroda T, Yanai H, Chuchottawon C, Kantima T, et al. Genomewide Association Study Confirming the Association of NAT2 with Susceptibility to Antituberculosis Drug-Induced Liver Injury in Thai Patients. Antimicrob Agents Chemother. 2019;63(8). doi: https://doi.org/10.1128/aac.02692-18
- Huerta-García AP, Medellín-Garibay SE, Salazar-González RA, Ortiz-Álvarez A, Magaña-Aquino M, Rodríguez-Pinal CJ, et al. Anthropometric and Genetic Factors Associated With the Exposure of Rifampicin and Isoniazid in Mexican Patients With Tuberculosis. Therapeutic Drug Monitoring. 2019;41(5):648-656. doi: https://doi.org/10.1097/ftd.0000000000000631
- Fredj NB, Romdhane HB, Woillard J, Chickaid M, Fadhel NB, Chadly Z, et al. Population pharmacokinetic model of isoniazid in patients with tuberculosis in Tunisia. International Journal of Infectious Diseases. 2021;104:562-567. doi: https://doi.org/10.1016/j.ijid.2021.01.033
- Zhao G, Chen M, Sun L, Xi N. Analysis of influencing factors on the plasma concentration of first-line anti-tuberculosis drugs-a single-center retrospective cohort study. Ann Transl Med. 2022;10(8):461-461. doi: https://doi.org/10.21037/atm-22-1341
- Thomas L, Raju AP, Chaithra, M SS, Varma M, Saravu K, et al. Influence of N-acetyltransferase 2 (NAT2) genotype/single nucleotide polymorphisms on clearance of isoniazid in tuberculosis patients: a systematic review of population pharmacokinetic models. Eur J Clin Pharmacol. 2022;78(10):1535-1553. doi: https://doi.org/10.1007/s00228-022-03362-7
- Ulanova V, Kivrane A, Viksna A, Pahirko L, Freimane L, Sadovska D, et al. Effect of NAT2, GSTM1 and CYP2E1 genetic polymorphisms on plasma concentration of isoniazid and its metabolites in patients with tuberculosis, and the assessment of exposure-response relationships. Front. Pharmacol. 2024;15. doi: https://doi.org/10.3389/fphar.2024.1332752
- Yunivita V, Gafar F, Santoso P, Chaidir L, Soeroto AY, Meirina TN, et al. Pharmacokinetics and pharmacodynamics of high-dose isoniazid for the treatment of rifampicin- or multidrug-resistant tuberculosis in Indonesia. Journal of Antimicrobial Chemotherapy. 2024;79(5):977-986. doi: https://doi.org/10.1093/jac/dkae057
- Rens NE, Uyl-de Groot CA, Goldhaber-Fiebert JD, Croda J, Andrews JR. Cost-effectiveness of a Pharmacogenomic Test for Stratified Isoniazid Dosing in Treatment of Active Tuberculosis. Clinical Infectious Diseases. 2020;71(12):3136-3143. doi: https://doi.org/10.1093/cid/ciz1212