RESEARCH ARTICLE
Phytochemical Analysis, In-vitro, and In-silico Antibacterial Activity of Stembark Extract of Anogeissus leiocarpus (DC) Guill and Perr
Sciences of Pharmacy|Vol. 2, Issue 3, pp. 134-147 (2023)
Received
Apr 10, 2023Revised
May 27, 2023Accepted
May 29, 2023Published
Jun 5, 2023
Abstract
Introduction
Bacterial infections (BI) coupled with antibiotic resistance (ABR) have been identified as the culprit leading to the death and morbidity of millions around the world, especially in places where there are a lot of challenges in the healthcare system such as developing countries (1). The menace of BI and ABR is expected to rise by 2050 with projected mortalities of up to 4 million worldwide (2, 3). In developing countries, poverty, poor access to modern healthcare facilities, and low government spending on healthcare might also contribute to the problems of BI and ABR (3-6). There are several antibacterial drugs available in the market but as earlier stated, poverty plays a role as these drugs are often expensive and unaffordable. Additionally, these drugs are unavailable to people living in rural areas and often associated with side effects. Furthermore, the quality of these drugs is questionable which may or may not contribute to the antibacterial resistance of the drugs. Thus, rural communities are forced to prospect for local sources of drugs to achieve therapeutic goals.
E. coli are leading cause of many bacterial infections in both humans and animals with notable infections including urinary tract infections, septicemia and enteritis (7). Additionally, neonatal meningitis is also caused by E. coli while in farm animals, diarrhea has been associated with E. coli. The antibiotics resistance E. coli to major classes of antibiotics such as β-lactams, quinolones, aminoglycosides third- and fourth-generation cephalosporins and monobactams attributed to its outer membrane barrier further complicating treatment (7). S. aureus are associated with different human infections notably bacteraemia, infective endocarditis, skin and soft tissue infections osteomyelitis, septic arthritis, prosthetic device infections, pulmonary infections gastroenteritis, meningitis, and urinary tract infections (8). The type and duration of the treatment depends on the type of infection, however, emergence of antibiotic resistance by this organism further complicates treatment (8). E. coli and S. aureus are two of the frequently encountered bacterial infections in humans with their treatment complicated by antibiotic resistance leading to prospects into alternative therapies such medicinal plants especially in developing countries.
Medicinal plants provide plant-based drugs which are stipulated as an alternative to antibacterial drugs, notably in low-income countries due to the availability, safety, and efficacy evidenced by the traditional use of the plants in traditional medicine (6, 9-11). The use of plant-based sources of drugs is often attributed to their phytoconstituents produced by plants for several purposes such as defense against pathogens other than growth and reproductive functions (12). The pharmacological actions of these plants are due to their individual and synergistic mode of action via restoration of normal body function by allowing healing to take place (13, 14). Medicinal plants exhibit several pharmacological effects including anti-inflammatory (15), and antimicrobial (16) effects via different mechanisms. The common antibacterial activity of medicinal plant extract is through the disruption bacterial of membrane functions, metabolic pathways, DNA and protein synthesis, and cell wall synthesis with synergistic mechanisms attributed to the inhibition of efflux pumps (16). The pharmacological properties of the medicinal plants are attributed to the phytochemical composition including alkaloids, flavonoids, and saponins.
Alkaloids exerts antibacterial effects via different mechanisms of action targeting different parts of bacteria and destroying its integrity. In a previous study, the alkaloids squalamine was reported to exert 16 to 32 times antibacterial effect than ciprofloxacin against Gram-negative pathogens (17). Indole-containing alkaloids were reported to exhibit antibacterial effects by inhibiting efflux pumps, the biofilm, filamentous temperature-sensitive protein Z, and methicillin-resistant Staphylococcus aureus pyruvate kinase (18). Furthermore, alkaloids were postulated to be novel sources of antibacterial therapeutic (19). Flavonoids are also attributed with antibacterial activities where in some cases exhibiting more potential than standard drugs against multi-drug resistant pathogens including Gram-negative and Gram-positive bacteria (20). (−)-Epigallocatechin was reported to exhibit antibacterial effect via DNA synthesis inhibition in Proteus vulgaris and RNA synthesis in S. aureus. Saponins were also reported to inhibit S. aureus in dose dependent manner with minimal MIC and MBC values. Specifically, quinoa saponins disrupted cell wall synthesis and degraded cytoplasmic and protein membranes leading to loss of cellular integrity (21).
Ethnobotanical surveys identified A. leiocarpus (AL) as a plant used in the traditional management of infections and diseases. The plant is often utilized as decoctions prepared by aqueous macerations taken orally to overcome an infection (22). In other cases, the plant parts are ground to powder and applied to external wounds to prevent infection to allow the healing process to take place (23). In experimental studies, AL was reported to exhibit antihyperglycemic (24), antioxidant (24), antihyperlipidemic (24), and antimicrobial effects (25) through different modes of action attributed to its phytoconstituents including alkaloids, glycosides, and flavonoids. However, there are limited studies revealing the potential mechanisms of action of the compounds present in the plants. Moreover, phytochemicals exhibit antibacterial effect through individual or synergistic mechanism of action targeting different molecules, proteins, and enzymes to exert their therapeutic effects which might counter antibiotic resistance (26, 27). Therefore, this study aimed to identify and quantitate the phytoconstituents of methanol stembark extract of A. leiocarpus and establish the antibacterial activity along with the potential mechanism of action in-silico as they are applied in folkloric medicine for antibacterial purposes.
Experimental Section
Materials
Collection of Plant Sample
AL was obtained from the Girei Local Government area, Adamawa State, Nigeria, followed by its identification and authentication by in the Forestry Department of Adamawa State Polytechnic, Yola where a voucher specimen was deposited (ASP/FT/245) after authentication by a Forest Technologist. It was cleaned, dried under shade, and ground to powder.
Reagents and Chemicals
Methanol, chloroform, ethyl acetate, butanol, and diethyl ether were purchased from Xingtai Dakun Technology Co. , Ltd (China). Nutrient agar and Mueller-Hinton agar (Qingdao Bio-Technology Co. , Ltd. , China). Amoxicillin (Zimilat®, Nemel Pharmaceutical, Nigeria). All other chemicals and reagents were of AnarlaR.
Methods
Phytochemicals Extraction and Analysis
The phytochemical extraction was done via 48 h maceration of 300 g of stem bark AL in 1 L of 70% v/v methanol, then filtered and dried over reduced pressure (28). The phytochemicals in methanol stembark extract of AL (MSEA) were detected by the standard method described previously as follow:
Alkaloids
The estimation of total alkaloids was done as previously described (29). Briefly, 0.5 g of the extract was weighed into a conical flask containing 10 mL of 10 % ammonium hydroxide to convert alkaloidal salts into the free base; the mixture was stirred and allowed to stand for 4 h before filtering. The filtrate was evaporated to one-quarter of its original volume on a water bath and concentrated ammonium hydroxide solution was added dropwise to the mixture to precipitate the alkaloids. The precipitate was filtered using a weighed filter paper and washed with 10% ammonium hydroxide solution. The precipitate was dried with the filter paper in an oven at 60°C for 30 minutes and then reweighed and calculated using Equation 1.
Where weight of residue = weight of the dried precipitate, weight of sample = weight of the extract taken earlier.
Saponins
Saponins were quantified by previously described methods (30). Exactly 0.5 g extract was introduced into a conical flask and 10 mL of 20% aqueous ethanol was added. The sample was heated over a water bath for 1 h with continuous stirring at about 550C. The concentrate was transferred into a 250 mL separator funnel and 5 mL of diethyl ether was added and shaken vigorously. The aqueous layer was recovered and the ether layer was discarded. About 10 mL of n-butanol was then added followed by the addition of 2 mL of 5% aqueous NaCl. The remaining solution was heated over a water bath. After evaporation, the sample was dried in the oven to a constant weight and calculated using Equation 1.
Flavonoids
Flavonoids were estimated by the previously described method (29). Briefly, 0.5g of the extract was mixed with 10 mL of 80% aqueous methanol. The whole solution was filtered through the Whatman filter paper. The filtrate was transferred to a pre-weighed crucible and evaporated into dryness over a water bath and weighed.
Antibacterial Activity
Bacterial Isolates Collection
The bacteria isolates were obtained from the microbiology laboratory of Modibbo Adama University Teaching Hospital, Yola, Nigeria which were subjected to characterization as previously described (31). Biochemical tests were carried out to ascertain isolated identity via the standard method (32, 33), followed by growing on nutrient agar and subsequent storage at 4 °C.
McFarland Standard (MS)
MS preparation was done by mixing 9.95 mL of 1% H2SO4 and 0.05 mL of 1.17% BaCl forming a precipitate that acted as a 0.5 MS turbidity for the isolates (33).
Inoculum Standardization
Inoculum was standardized by culturing on nutrient agar and incubated at 37 °C overnight followed by transferring the formed colonies to test tubes with 5 mL of 0.9% normal saline adjusted to the turbidity of the MS (34).
Zone of Inhibition (ZI)
To ascertain the antibacterial activity of AL, a slightly modified agar diffusion technique was applied (35). The isolates were inoculated on a solidified Mueller-Hinton (MH) agar, followed by the addition of 0.2 mL extract at varied concentrations and added to five wells with the sixth acting as a positive control containing amoxicillin at 50 mg/mL. The mixture was incubated overnight at 37 °C. The antibacterial effect of the extract was expressed by the diameter of the ZI in mm.
Minimum Inhibitory Concentration (MIC)
The MIC was evaluated according to the protocols of the National Committee for Clinical Laboratory Standards (NCCLS) (36). One milliliter of extract was dispensed into 5 mL of MH broth containing test tubes and mixed to which 0.1 mL of the isolate broths were added and incubated overnight at 37 °C. The minimum concentration at which the bacterial growth was completely inhibited was defined as the MIC of the extracts.
Bactericidal Concentration (MBC)
Further evaluation of MBC was done by subculturing the test tube without visible growth in the MIC and incubating overnight at 37 °C. The least concentration without visible bacteria growth defined the MBC (37).
Molecular Docking
The compounds used for the in-silico study were collected from our previous study (38) were downloaded from PubChem (https://pubchem. ncbi. nlm. nih. gov) along with the target inhibitors while the antibacterial targets were downloaded from the RSCB protein data bank (https://www. rcsb. org) in PDB format. PubChem and RSCB IDs for compounds, inhibitors, and targets were documented. Table 1 lists compound and standard drug PubChem IDs, while Table 2 details enzyme targets with PDB ID, grid coordinates, and box size.
| Compounds | ||
|---|---|---|
| Name | PubChem ID | Designation |
| 5-Methyl-1H-pyrazole-3-carboxylic acid | 9822 | Compound I |
| 1 2 4-Benzenetriol | 10787 | Compound II |
| 2-Methoxyhydroquinone | 69988 | Compound III |
| Maltol | 8369 | Compound IV |
| Methyl 14-methylpentadecanoate | 21205 | Compound V |
| 5-Hydroxymethylfurfural | 237332 | Compound VI |
| 1 2-Epoxyhexadecane | 23741 | Compound VII |
| 2-(Tetradecyloxy) ethanol | 16491 | Compound VIII |
| Methyl palmitate | 8181 | Compound IX |
| Hexadecanal | 984 | Compound X |
| Standard Drugs | ||
| Sulfamethoxazole | 5329 | Compound INB1 |
| Cefepime | 5479537 | Compound INB2 |
| Methotrexate | 126941 | Compound INB3 |
| Fosfomycin | 446987 | Compound INB4 |
| Ciprofloxacin | 2764 | Compound INB5 |
| Afabicin | 72696796 | Compound INB6 |
| Name | PDB ID | Grid coordinate | Box size |
|---|---|---|---|
| Dihydropteroate synthase | 5V79 (39) | X=-21.57, Y=9.40, Z=102.75 | X=25.52, Y=18.36, Z=22.69 |
| Penicillin-binding protein 2X | 5OJ0 (40) | X=33.90, Y=-16.78, Z=54.66 | X=24.15, Y=23.50, Z= 18.11 |
| Dihydrofolate reductase | 1RG7 (41) | X=-0.97, Y=21.43, Z=21.19 | X=21.18, Y= 18.56, Z=22.96 |
| MurA | 2RL2 (42) | X=25.60, Y=-49.61, Z=51.90 | X=20.25, Y=24.06 Z= 19.76 |
| Topoisomerase IV | 3FV5 (43) | X=13.379, Y=-0.70, Z=2.53 | X= 23.55, Y= 17.65, Z=19.11 |
| FabI | 1LX6 (44) | X=-2.01, Y=22.40, Z=134.49 | X=15.62, Y=17.70, Z=17.90 |
The protein/receptor targets used for the present study were selected because they are targets of different antibacterial drugs (the roles of the proteins can be seen in the discussion section). The protein targets were downloaded already docked with their inhibitors prior to preparation using AutoDock Tools. The amino acid residues interacting with the inhibitors were marked and subsequently selected while choosing grid coordinate and box size.
Preparation of the Compounds, Inhibitors, and Targets
The list of the compounds along with their PubChem ID is provided in Table 1 while the targets and inhibitors with their PDB ID and PubChem ID respectively are provided in Table 2. The compounds and inhibitors were downloaded and converted to PDB format with Openbabel software version 3.1.1 (45). The targets downloaded in PDB format were prepared using AutoDock Tools, removing water molecules and hetero atoms 1.5.7 (46) and saved in PDB format to allow for proper docking of the ligands (compounds and inhibitors) with the target. The compounds and inhibitors downloaded were further subjected to energy minimization using the PyRx 0.8 software before docking.
Docking Procedure
The virtual screening of all the compounds and inhibitors against all the the targets was carried out using PyRx 0.8 software via the vina wizard with the exhaustiveness set 32. The ligand with lowest binding energy was selected and saved as PDB for visualization using PyMOL 2.5.4 and 2D visualization was done with LigPlot+ 2.2.8. The profiler web server was utilized to visualize other binding interactions (47). The binding energy was used to determine the inhibition constant (Ki) using the formula Ki = exp ∆G/RT, where ∆G is the binding energy, R is the universal gas constant = 1.985 x 10-3 kcal-1 mol-1 k-1, and T is the temperature (298.15 K) (48).


Statistics
The values obtained were expressed as mean ± standard error of triplicate determinations' mean (± SEM) and evaluated with Statistical Package for the Social Sciences (SPSS) version 22 software. One-way analysis of variance was used to assess the differences among the groups means followed by the Tukey multiple comparison test at p< 0.05.
Result
The phytochemical components identified in the AL extract were alkaloids, saponins, and flavonoids. Alkaloids had the least concentration of 7.17 ± 0.60%, while saponins had a concentration of 11.33 ± 3.18%. Flavonoids were quantified in the highest concentration of 31.01 ± 4.04%. Figure 1 shows the antibacterial effects demonstrated by the AL extract on the bacteria isolates revealed by the ZI. A maximum ZI of 13.5 ±1.21 mm and 9 ±1.02 mm was observed for E. coli and S. aureus respectively at 100 mg/mL concentration. The standard amoxicillin exhibited a ZI of 30 ±1.00 mm and 20 ±2.00 mm for E. coli and S. aureus respectively at the concentration of 50 mg/mL. The least inhibitory effect was observed to be 7.0 ±1.21 mm and 2.0 ±1.31 mm respectively for E. coli and S. aureus at 25 mg/ml concentration.
The inhibitory effects of AL extract are presented in Table 4. The MIC of the AL extract against E. coli and S. aureus were 12.5 mg/mL and ≤ 6.25 mg/mL respectively. Table 4 also presents the findings of the MBC which is the lowest concentration at which the bacterial isolates are neutralized by the (AL) extract. At a concentration > 100 mg/mL E. coli was not neutralized while S. aureus was completely neutralized at 100 mg/mL.
| Test | Test organism | Incubation time (Hours) | Concentration (mg/ml) | Remark | ||||
|---|---|---|---|---|---|---|---|---|
| 100 | 50 | 25 | 12.5 | 6.25 | ||||
| MIC | E. coli | 24 | − | − | − | − | + | 12.5 |
| S. aureus | 24 | − | − | − | − | − | ≤ 6.25 | |
| MBC | E. coli | 24 | + | + | + | + | + | > 100 |
| S. aureus | 24 | − | + | + | + | + | 100 | |
Supplemental Table 1 (Sheet 1) provides an overview of the docking interactions between dihydropteroate synthase (DS) and various compounds, including the inhibitor sulfamethoxazole (INB1). Among the compounds, Compound IX (methyl palmitate) demonstrated the highest ranking based on binding affinity (BA) (-6.4 kcal/mol) and inhibition constant (Ki) (20 µM), along with INB1. While both INB1 and Compound IX shared similar BA and Ki values, INB1 exhibited a greater number of hydrogen bond (HB) interactions (7). On the other hand, Compound VIII (2-(Tetradecyloxy) ethanol) had the highest BA (-4.2 kcal/mol) and Ki (828 µM), indicating it as the least favorable interaction with DS. Among the compounds, Compound II (1, 2, 4-Benzenetriol) displayed a specific cation interaction (CI) with Arg255, which was the only such interaction observed. To visually illustrate the docking interactions, Figure 2 presents both the 2D and 3D representations of Compound IX and INB1, highlighting the amino acids involved in hydrophobic interactions (HBI) and HB, with the corresponding distances measured in angstrom units.
The molecular docking interactions of penicillin-binding protein 2X (PBP 2X) with the compounds and its inhibitor are presented in Supplemental Table 1 (Sheet 2). The best docking pose with the lowest BA (-7.9 kcal/mol) and Ki (2 µM) was exhibited by INB2 (Cefepime) with 1 HB, 12 hydrophobic interactions (HBI). Compound IX (methyl palmitate) had the lowest BA (-7.1 kcal/mol) and Ki (6 µM) among the compounds while compound VII (1, 2-Epoxyhexadecane) had the highest BA (-4.4 kcal/mol) and Ki (590 µM) with 2 HB and 7 HBI. Only compound I (-Methyl-1H-pyrazole-3-carboxylic acid) demonstrated salt bridge interaction among the compounds by interacting with Lys340. Figure 3 shows the amino acids involved in the interactions of compound II and INB2 with PBP 2X with accompanied HBIs including the HB distance in angstrom.
The docking interactions between dihydrofolate reductase (DS) and various compounds, along with its inhibitor, are presented in Supplemental Table 1 (Sheet 3). INB3 (Methotrexate) demonstrated the lowest BA (-9.7 kcal/mol) and Ki (7.6 × 10-2 µM), along with 9 hydrogen bond (HB) interactions, 12 hydrophobic interactions (HBI), 1 cation interaction (CI), and 2 salt bridge (SB) interactions, positioning it as the most favorable docking pose. Compound VIII (2-(Tetradecyloxy) ethanol) exhibited the lowest BA (-6.2 kcal/mol) and Ki (28 µM) among the compounds, featuring 3 HB interactions and 15 HBI interactions. Notably, only compounds V and IX formed salt bridges with Arg255 among the compounds. Figure 4 visualizes the 2D and 3D interactions of dihydrofolate reductase (DS), highlighting the similarities in amino acid interactions between compound VIII and INB3.


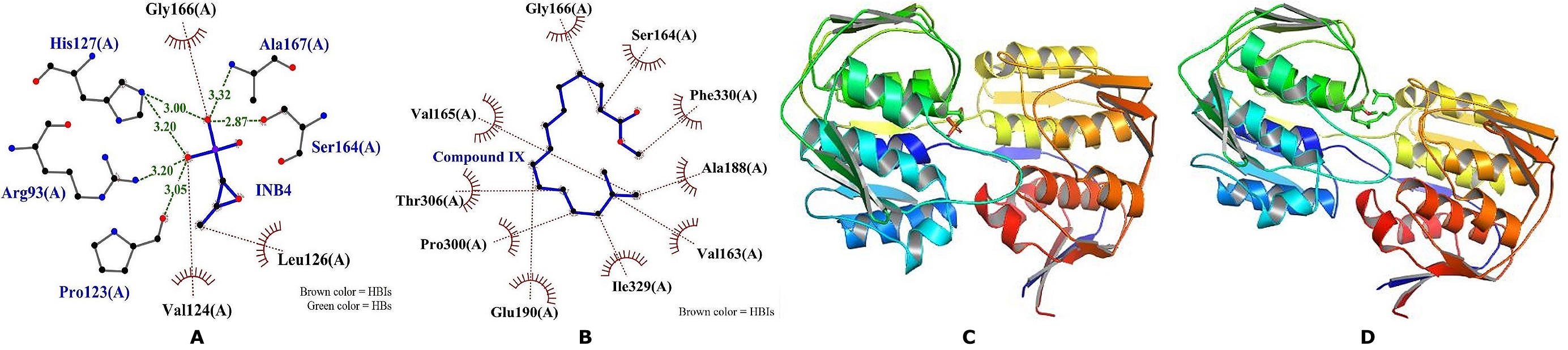


The interactions between UDP-N-acetylglucosamine enolpyruvyl transferase (Mur A) and its inhibitor, along with the compounds demonstrating their binding affinity (BA) and inhibition constant (Ki), are presented in Supplemental Table 1 (Sheet 4). Among the compounds, Compound IX (methyl palmitate) displayed the lowest BA (-7.2 kcal/mol) and Ki (5 µM), surpassing all other compounds, including INB4 (Fosfomycin). Interestingly, all the other compounds exhibited lower BA and Ki values than INB4, with Compound VI showing the least favorable interaction, featuring a BA of -4.4 kcal/mol and Ki of 590 µM. Compounds IV and II also engaged in specific cation interactions with Arg93 and Arg333, respectively. Figure 5 illustrates the amino acids involved in the interactions between MurA and INB5, as well as Compound IX, highlighting the hydrogen bond (HB) and hydrophobic interaction (HBI) patterns, along with the corresponding HB distances measured in angstrom units.
Supplemental Table 1 (Sheet 5) provides an overview of the docking interactions between topoisomerase IV (TopoIV) and its inhibitor, as well as the compounds, showcasing their binding affinity (BA), inhibition constant (Ki), hydrogen bond (HB) interactions, hydrophobic interactions (HBI), and van der Waals interactions (VWI). INB5 (Ciprofloxacin) displayed the lowest BA (-7.8 kcal/mol) and Ki (2 µM), accompanied by 3 HB interactions, 7 HBI interactions, and a salt bridge linkage with Glu46. Among the compounds, Compound VI (5-Hydroxymethylfurfural) and I (5-Methyl-1H-pyrazole-3-carboxylic acid) demonstrated the lowest BA (-5 kcal/mol) and Ki (214 µM). However, Compound VI exhibited 4 HB interactions and 4 HBI interactions, while Compound III (2-Methoxyhydroquinone) showcased the highest BA (-4.2 kcal/mol) and Ki (828 µM) with 1 HB interaction and HBI. Figure 6 visually presents the 2D and 3D binding interactions of INB6 and Compound VI, including the amino acids involved and the corresponding HB distances.
Figure 7 provides a visual representation of the 2D and 3D interactions between INB6, Compound IX, and FabI, highlighting the HB interactions, HBI interactions, and the corresponding HB distances measured in angstrom units. The docking interactions between Enoyl-acyl-carrier-protein Reductase (FabI) and various compounds, including INB6 (Afabicin), are outlined in Supplemental Table 1 (Sheet 6). INB6 demonstrated the lowest binding affinity (BA) (-7.6 kcal/mol) and inhibition constant (Ki) (3 µM), with 2 hydrogen bond (HB) interactions and 8 hydrophobic interactions (HBI). Among the compounds, Compound IX (methyl palmitate) exhibited the least BA (-5.8 kcal/mol) and Ki (3 µM), featuring 7 HBI interactions without any HB interactions. On the other hand, Compound II (1, 2, 4-Benzenetriol) displayed the highest BA (-3.7 kcal/mol) and Ki (1927 µM), accompanied by a higher number of HB interactions compared to Compound IX. Notably, none of the compounds, including the inhibitor, showed any cation interaction (CI), salt bridge (SB), or pi-stacking (PS) interactions.
Table 5 summarizes the BA and Ki of the compounds and inhibitors of the target enzymes. Among the compounds, III (2-Methoxyhydroquinone) exhibited the least BA and Ki against 3 out of the 5 enzymes demonstrating a BA (-5.7 kcal/mol) against PBP2x, the least among all the compounds across all the targets, thus, might be partly responsible for the antibacterial action of AL.
| Enzymes | Compound | Standard Inhibitors | ||||
|---|---|---|---|---|---|---|
| Designation | BA (kcal/mol) | Ki (µM) | Designation | BA (kcal/mol) | Ki (µM) | |
| DS | IX | -6.4 | 20 | INB1 | -6.4 | 20 |
| PBPX2 | IX | -7.1 | 6 | INB2 | -7.9 | 2 |
| DR | VIII | -6.2 | 28 | INB3 | -9.7 | 7.6 × 10-2 |
| Mur A | IX | -7.2 | 5 | INB4 | -4.5 | 499 |
| TopoIV | VI | -5 | 214 | INB5 | -7.8 | 2 |
| FabI | IX | -5.8 | 55 | INB6 | 7.6 | 3 |
Discussion
Phytochemicals which are secondary metabolites including alkaloids produced by plants were reported to exert antibacterial effects with broad-spectrum effects (17, 19). The antibiotic-enhancing and anti-virulence activity of flavonoids was previously reported (17). Indole alkaloids isolated from Pseudomonas aeruginosa were reported to exhibit potent antimicrobial action towards gram-negative and positive bacteria (49). Saponin compounds isolated from Chenopodium quinoa demonstrated anti-bactericidal activity towards S. aureus, S. epidermidis, and B. cereus with the highest activity recorded against S. aureus (21). Saponins from Albizia adianthifolia exerted considerable antibacterial effects against multi-drug resistant gram-negative bacteria (50). Flavonoids exert an antibacterial effect via disruption of the cell wall, protein, nucleic acid synthesis, and energy metabolism (51). Additionally, the cell membrane was predicted to be the target of flavonoids via phospholipid bilayer damage and disruption of ATP synthesis (52). Flavonoid reported in our study was not detected in a previous study on the methanol partitioned extract of AL (53). Similarly in another study, flavonoids were absent (54). In another study, alkaloids were absent in the MSEA but saponins and flavonoids were observed (55). The variation in the detection of phytochemicals in the methanol AL extract might be attributed to the difference in extraction methods (56).
The antibacterial activity of Anogeissus lieocarpus, against the test bacteria, justified its traditional use as an ethnomedicinal plant by locals in the study area. The presence of the identified phytochemicals may warrant the plant's inhibitory function. Amoxicillin, the positive control had the widest zone of inhibition. Compared to the crude extracts, this antibiotic demonstrated higher activity. This is not astonishing because it is expected that standard antibiotics should exert superior activity due to their refined natured compared to crude extracts. The relatively thin peptidoglycan layer of gram-negative bacteria and an outer phospholipidic membrane contain lipopolysaccharide components that result in lipophilic solute impermeability for Gram-positive bacteria, outer peptidoglycan layers are thick thus, not an effective and excellent permeable barrier which increase susceptibility to the plant extract (57). The MIC spanned from 6.25 to 12.5 mg/mL which is lower than the values previously documented (54, 58), attributed to the presence of active phytochemical constituents which inhibits bacterial growths. A bactericidal activity of AL extract was observed at 100 mg/ml for S. aureus but no impact on E. coli showing the relative effect of concentrations, thus suggesting greater concentrations may be necessary for E. coli as the present concentration maybe bacteriostatic. Nonetheless, published results on the antibacterial effect of AL reported the lethal effects of the extract on S. aureus (59) agreeing with the present study. The MIC values in this study were different and lower than the MBC value obtained demonstrating that rather than killing the organisms, the concentration used in the study was only able to inhibit their growth. This is in tandem with the study carried out previously (60). The lower MIC value recorded for the study provides proof of the potent antibacterial properties of A, consistent and in tandem with a previous study demonstrating a variety of actions against a wide range of bacterial pathogens (61).
Dihydropteroate synthase (DS) catalyzes the synthesis of dihydropteroate via the conversion of 6-hydroxymethyl-dihydropterin 1′-diphosphate and 4-aminobenzoate yielding dihydropteroate and inorganic pyrophosphate. This enzyme act as a target for antimicrobial drugs such as Sulfamethoxazole because folic acid is not synthesized by humans who depend on preformed folic acid. Sulfamethoxazole is an analog of para-aminobenzoic acid (PABA) acting by competitively binding to the enzyme preventing the binding of PABA which is the substrate for folic acid synthesis (62). Although compound IX interacted with higher BA and Ki, both compounds interacted with a similar amino acid including Arg255, Lys221, Thr62, and Phe190 with compound IX exhibiting HBI with all the amino acids while INB1 exhibited HB with Thr62 and Phe98. The hydrophobic nature of compound IX might contribute to the favorable interaction of compound IX with the DS binding pocket compared to the other compounds. Although both INB1 and compound IX displayed the same BA and Ki, the superior HB demonstrated by INB1 might offer a stronger interaction with DS binding pocket than compound IX and offers more stability for the complex formed. The binding of compound IX to DS might disrupt the activity of the enzyme preventing folic acid synthesis via competitive inhibition. Thus, the antibacterial effect of the AL extract might be partly attributed to the action of compound IX, though a further study of the binding site might be required to justify this claim. The fungicidal effects of compound IX and its derivatives were previously reported (63). Additionally, derivatives of compound II (1, 2, 4-Benzenetriol), were reported to exert antibacterial activities depicting the compound as a precursor for novel antibacterial compounds (64).
Penicillin-binding protein 2X (PBP 2X) is a class B PBP anchored to the membrane participating in the final stage of peptidoglycan synthesis making them targets of antibiotics specifically the β-lactam (65). β-lactam like INB2 (Cefepime) exhibits suicide inhibition covalently binding to the active site of PBP 2X via acylation for an extended period forming in active complex and preventing the catalytic action of the enzyme subsequently blocking peptidoglycan synthesis and causing cell lysis (66). Compound IX with the least BA and Ki among the compounds exhibited a slightly higher BA and Ki compared to INB2 interacting with similar amino acids though INB2 formed HB with Asn337. The low BA and Ki of compound IX for PBP 2X might contribute to the antibacterial effect of AL preventing the catalytic effect of the PBP 2X via covalent bonding. Although Compound VI exhibited higher BA compared to compound IX and INB2, it demonstrated a higher number of HB which might translate to better stability. Thr550, Ser395, Ser337, Asn397, Glu552, Gln452, and Lys340 were the identified amino acids interacting with INB2 and compound IX participating in different binding interactions.
Dihydrofolate reductase (DR) catalyzes the formation of 5, 6, 7, 8-tetrahydrofolic acid via the reduction of 7, 8-dihydrofolate through hydride transfer from the cofactor NADPH to the pterin ring yielding 5, 6, 7, 8-tetrahydrofolic acid and NADP (67). This is a critical reaction for maintaining tetrahydro folic acid level required for nucleotide synthesis required for cell growth and proliferation, thus a major target of antibacterial drugs considering the rapidly dividing nature of bacterial cells (67). Methotrexate exerts a broad-spectrum antibacterial effect against gram-positive and some gram-negative by binding to DR thereby inhibiting folic acid synthesis required for cell growth and proliferation (68). In our study, INB3 (Methotrexate) exhibited a superior binding to DR than all the compounds forming cation interactions with Phe31 and salt bridge with Asp27 and Lys32 in addition to the 9 HB and 12 HBI. This creates a better and more stable interaction with the enzyme and inhibits its activity. Compound VIII showed the least BA and Ki among the compounds that exhibited fewer HB but more HBI than INB3. Some of the key amino acids identified participating in binding interactions of DR with INB3 and compound VIII include Arg57, Asp27, Ile5, and Ile94 which might be crucial for inhibiting the activity of DR, thus antibiotic targets.
MurA enzyme is a transferase that catalyzes the transfer of the enolypyruvate moiety of phosphoe-nolpyruvate (PEP) to UDP-n-acetylglucosamine (UDP-GlcNAc) which marks the first step of the Mur pathway during bacteria cell wall synthesis and being absent in eukaryotes makes them targets of many antibiotics (69). Fosfomycin interferes with bacterial cell wall synthesis by inhibiting the action of MurA via inactivating the enzyme by covalently binding to the active site of the enzyme preventing the early cytoplasmic stage of the cell wall synthesis (70). In the present study, INB4 exhibited higher BA and Ki against MurA compared to all the compounds except compound VI. This might be attributed to the non-interaction of fosfomycin with the key active site residues (Cys117 and Ser118) even though none of the compounds also interacted with these amino acids. Compound IX (methyl palmitate) exhibited the lowest BA and Ki among all the compounds which might be attributed to the hydrophobic nature of the compounds contributing to its stable interactions with residues within binding pockets.
Topoisomerase IV (TopoIV) is critical in maintaining the viability and genetic stability of cells by unraveling the newly formed DNA during replication to allow for the daughter chromosome separation as both the replication and segregation occur concurrently during cell division (71). TopoIV serves as an ideal target for many antibiotics inhibiting cell division such as ciprofloxacin which acts via topoisomerase II and IV inhibition (72). Ciprofloxacin is a broad-spectrum antibiotic for gram-negative and positive bacteria, binding its microbial target with 100 times more affinity than the mammalian target (73). In our study, INB5 (ciprofloxacin) exhibited superior BA and Ki compared to all the compounds with additional SB formation with Glu46. Among the compounds, VI (5-Hydroxymethylfurfural) and I (5-Methyl-1H-pyrazole-3-carboxylic acid) exhibited superior BA and Ki with compound VI showing a more stable interaction with the enzyme with more HB. Compound VI interacted with similar residues with INB5 but formed more HB which might be translated to extended binding time and lasting effect on the enzyme than INB5. The binding of compound IV to TopoIV with stability might disrupt the activity of the enzyme with bactericidal effects. Compound IV was previously linked to antibacterial activities (74). In another study, the compound was attributed with antibacterial effects against Acinetobacter baumanni through inhibition of biofilm formation and suppression of virulence regulator genes (75).
The fabI gene encodes the fabI reductase enzyme, a rate-limiting enzyme in the FAS-II pathway and an NADH-dependent enzyme catalyzing the last reaction of each round of elongation during the reduction of an enoyl-acyl carrier protein, thus a broad-spectrum antibacterial target and development of novel antibiotics (76). Afabicin is a first-class antibiotic targeting the bacterial fatty acid synthesis pathway (FAS-II) inhibiting the action of enoyl-acyl carrier protein reductase (FabI) (77). In the present study, INB6 demonstrated the least BA and Ki compared to all the other compounds displaying superior inhibitory effects against fabI forming HB with Gly93 and Lys163 along with 8 HBI. This might contribute to the stability of the complex formed with an extended binding period. Among the compounds, IX exhibited the least BA and Ki without HB interactions which might be attributed to the hydrophobic nature of the compound
Conclusion
The present study evaluated the antibacterial actions of AL for its acclaimed use in folkloric medicine. AL demonstrated antibacterial activity evidenced by the bacterial growth inhibition and bactericidal potential displayed by the plant in-vitro which might be attributed to the presence of phytochemicals. Furthermore, the in-silico study scientifically justifies the use of the effectiveness of the plant in the treatment of bacterial infections as claimed in folkloric medicine.
Declarations
Acknowledgment
The authors would like to thank the Department of Science Laboratory Technology, Adamawa State Polytechnic Yola, for their institutional support.
Conflict of Interest
The authors declare no conflicting interest.
Data Availability
You can access the Supplemental Table 1 on the ETFLIN server via the following link: https://etflin.com/file/document/202305270733301064215839.xlsx
Ethics Statement
Not applicable.
Funding Information
Not applicable.
References
- Church NA, McKillip JL. Antibiotic resistance crisis: challenges and imperatives. Biologia. 2021;76(5):1535-50.
- Elton L, Thomason MJ, Tembo J, Velavan TP, Pallerla SR, Arruda LB, et al. Antimicrobial resistance preparedness in sub-Saharan African countries. Antimicrobial Resistance & Infection Control. 2020;9:1-11.
- Akinde O, Taiwo M. Emerging antibiotic resistance in africa; threat to healthcare delivery. MOJ Biology and Medicine. 2017;1(4):114-115.
- Gandra S, Alvarez-Uria G, Turner P, Joshi J, Limmathurotsakul D, van Doorn HR. Antimicrobial Resistance Surveillance in Low- and Middle-Income Countries: Progress and Challenges in Eight South Asian and Southeast Asian Countries. Clin Microbiol Rev. 2020;33(3):e00048-19.
- Dunachie SJ, Day NPJ, Dolecek C. The challenges of estimating the human global burden of disease of antimicrobial resistant bacteria. Curr Opin Microbiol. 2020;57:95-101.
- Sartelli M, C. Hardcastle T, Catena F, Chichom-Mefire A, Coccolini F, Dhingra S, et al. Antibiotic Use in Low and Middle-Income Countries and the Challenges of Antimicrobial Resistance in Surgery. Antibiotics [Internet]. 2020; 9(8):1-12.
- Allocati N, Masulli M, Alexeyev MF, Di Ilio C. Escherichia coli in Europe: An Overview. International Journal of Environmental Research and Public Health [Internet]. 2013; 10(12): 235-254. Available from: http://dx.doi.org/10.3390/ijerph10126235
- Tong SYC, Davis JS, Eichenberger E, Holland TL, Fowler Jr VG. Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clin Microbiol Rev. 2015;28(3):603-667
- Süntar I. Importance of ethnopharmacological studies in drug discovery: role of medicinal plants. Phytochemistry Reviews. 2020;19(5):1199-209.
- Kaggwa B, Kyeyune H, Munanura EI, Anywar G, Lutoti S, Aber J, et al. Safety and Efficacy of Medicinal Plants Used to Manufacture Herbal Products with Regulatory Approval in Uganda: A Cross-Sectional Study. Evid Based Complement Alternat Med. 2022;2022:1304839.
- Deyno S, Eneyew K, Seyfe S, Wondim E. Efficacy, safety and phytochemistry of medicinal plants used for the management of diabetes mellitus in Ethiopia: a systematic review. Clinical Phytoscience. 2021;7(1): 1-16.
- Kaushik B, Sharma J, Kumar P, Shourie A. Phytochemical properties and pharmacological role of plants: secondary metabolites. Biosciences Biotechnology Research Asia. 2021;18(1):1-23.
- Singh V. Medicinal plants and bone healing. National journal of maxillofacial surgery. 2017;8(1):1-8
- Ugboko HU, Nwinyi OC, Oranusi SU, Fatoki TH, Omonhinmin CA. Antimicrobial importance of medicinal plants in Nigeria. The Scientific World Journal. 2020;2020: 1-10.
- Bouyahya A, Guaouguaou F-E, El Omari N, El Menyiy N, Balahbib A, El-Shazly M, et al. Anti-inflammatory and analgesic properties of Moroccan medicinal plants: Phytochemistry, in vitro and in vivo investigations, mechanism insights, clinical evidences and perspectives. Journal of Pharmaceutical Analysis. 2022;12(1):35-57.
- Álvarez-Martínez FJ, Barrajón-Catalán E, Herranz-López M, Micol V. Antibacterial plant compounds, extracts and essential oils: An updated review on their effects and putative mechanisms of action. Phytomedicine. 2021;90:153626.
- Cushnie TPT, Cushnie B, Lamb AJ. Alkaloids: An overview of their antibacterial, antibiotic-enhancing and antivirulence activities. Int J Antimicrob Agents. 2014;44(5):377-386.
- Liu Y, Cui Y, Lu L, Gong Y, Han W, Piao G. Natural indole-containing alkaloids and their antibacterial activities. Arch Pharm. 2020;353(10):2000120.
- Yan Y, Li X, Zhang C, Lv L, Gao B, Li M. Research Progress on Antibacterial Activities and Mechanisms of Natural Alkaloids: A Review. Antibiotics [Internet]. 2021; 10(3). 318. Available from: http://dx.doi.org/10.3390/antibiotics10030318
- Farhadi F, Khameneh B, Iranshahi M, Iranshahy M. Antibacterial activity of flavonoids and their structure–activity relationship: An update review. Phytother Res. 2019;33(1):13-40.
- Dong S, Yang X, Zhao L, Zhang F, Hou Z, Xue P. Antibacterial activity and mechanism of action saponins from Chenopodium quinoa Willd. husks against foodborne pathogenic bacteria. Industrial Crops and Products. 2020;149:112350.
- Sani S, Aliyu B, Haruna M, Yahya SM, Yakasai MA, Hayatu LW, et al. Local plants and diabetes management; folkloric practices in Metropolitan Kano, Nigeria. Bayero Journal of Pure and Applied Sciences. 2019;12(1):268-273.
- Kankara SS, Ibrahim MH, Mustafa M, Go R. Ethnobotanical survey of medicinal plants used for traditional maternal healthcare in Katsina state, Nigeria. South African Journal of Botany. 2015;97:165-175.
- Motto AE, Lawson-Evi P, Eklu-Gadegbeku K. Antidiabetic and antioxidant potential of total extract and supernatant fraction of the roots of Anogeissus leiocarpus in HFD-fed and Streptozocin -induced diabetic rats. Biomed Pharmacother. 2022;154:113578.
- Alhassan DA, Uba AI, Muhammad AU, Muhammad YYu. Phytochemical screening and antimicrobial activity of crude stem bark extracts of Anogeissus leiocarpus. European Journal of Medicinal Plants. 2016;11(2):1-7.
- Ayaz M, Ullah F, Sadiq A, Ullah F, Ovais M, Ahmed J, et al. Synergistic interactions of phytochemicals with antimicrobial agents: Potential strategy to counteract drug resistance. Chem Biol Interact. 2019;308:294-303.
- Carvalho RS, Carollo CA, de Magalhães JC, Palumbo JMC, Boaretto AG, Nunes e Sá IC, et al. Antibacterial and antifungal activities of phenolic compound-enriched ethyl acetate fraction from Cochlospermum regium (mart. Et. Schr.) Pilger roots: Mechanisms of action and synergism with tannin and gallic acid. South African Journal of Botany. 2018;114:181-187.
- Evans WC. Trease and Evans' pharmacognosy: Elsevier Health Sciences; 2009.
- Harborne A. Phytochemical methods a guide to modern techniques of plant analysis: springer science & business media; 1998.
- Obadoni B, Ochuko P. Phytochemical studies and comparative efficacy of the crude extracts of some haemostatic plants in Edo and Delta States of Nigeria. Global Journal of pure and applied sciences. 2002;8(2):203-8.
- Abubakar I, Usman A. Phytochemical and antibacterial investigations of moringa (Moringa oleifera) leaf extract on selected bacterial pathogens. 2016. Journal of Microbiology and Antimicrobials. 2016;8(5):28-33.
- Talaiekhozani A. Guidelines for quick application of biochemical tests to identify unknown bacteria. Account of Biotechnology Research (2013). Available at http://dx.doi.org/10.2139/ssrn.4101035
- Cheesbrough M. Biochemical tests to identify bacteria. District laboratory practice in tropical countries, Part. 2002;2:63-70.
- Andrews JM. BSAC standardized disc susceptibility testing method (version 4). J Antimicrob Chemother. 2005;56(1):60-76.
- Biradar YS, Jagatap S, Khandelwal KR, Singhania SS. Exploring of antimicrobial activity of Triphala Mashi—An ayurvedic formulation. Evid Based Complement Alternat Med. 2008;5(1):107-113.
- Lar PM, Ojile EE, Dashe E, Oluoma JN. Antibacterial Activity on Moringa Oleifera Seed Extracts on Some Gram-Negative Bacterial Isolates. African Journal of Natural Sciences. 2011;4:57-62.
- De NB, Ifeoma E. Antimicrobial effects of components of the bark extract of neem (Azadirachta indica A. J uss). Technology and Development. 2002;8:23-28.
- Dahiru MM, Badgal EB, Neksumi M. Phytochemical profiling and heavy metals composition of aqueous and ethanol extracts of Anogeissus leiocarpus. Journal of Faculty of Pharmacy of Ankara University. 2023;47(2):311-323.
- Dennis ML, Lee MD, Harjani JR, Ahmed M, DeBono AJ, Pitcher NP, et al. 8‐Mercaptoguanine Derivatives as Inhibitors of Dihydropteroate Synthase. Chemistry–A European Journal. 2018;24(8):1922-1930.
- Bernardo-García N, Mahasenan KV, Batuecas MT, Lee M, Hesek D, Petrackova D, et al. Allostery, recognition of nascent peptidoglycan, and cross-linking of the cell wall by the essential penicillin-binding protein 2x of Streptococcus pneumoniae. ACS Chem Biol. 2018;13(3):694-702.
- Sawaya MR, Kraut J. Loop and subdomain movements in the mechanism of Escherichia coli dihydrofolate reductase: crystallographic evidence. Biochemistry. 1997;36(3):586-603.
- Yoon HJ, Lee SJ, Mikami B, Park HJ, Yoo J, Suh SW. Crystal structure of UDP‐N‐acetylglucosamine enolpyruvyl transferase from Haemophilus influenzae in complex with UDP‐N‐acetylglucosamine and fosfomycin. Proteins: Structure, Function, and Bioinformatics. 2008;71(2):1032-1039.
- Wei Y, Charifson P, LeTiran A. Crystal Structure of E. coli Topoisomerase IV co-complexed with inhibitor. 2010. Available from: https://doi.org/10.2210/pdb3fv5/pdb.
- Miller WH, Seefeld MA, Newlander KA, Uzinskas IN, Burgess WJ, Heerding DA, et al. Discovery of aminopyridine-based inhibitors of bacterial enoyl-ACP reductase (FabI). J Med Chem. 2002;45(15):3246-56.
- O'Boyle NM, Banck M, James CA, Morley C, Vandermeersch T, Hutchison GR. Open Babel: An open chemical toolbox. Journal of cheminformatics. 2011;3(1):1-14.
- Sanner MF. Python: a programming language for software integration and development. J Mol Graph Model. 1999;17(1):57-61.
- Adasme MF, Linnemann KL, Bolz SN, Kaiser F, Salentin S, Haupt VJ, et al. PLIP 2021: Expanding the scope of the protein–ligand interaction profiler to DNA and RNA. Nucleic Acids Res. 2021;49(W1):W530-W534.
- Ortiz CLD, Completo GC, Nacario RC, Nellas RB. Potential Inhibitors of Galactofuranosyltransferase 2 (GlfT2): Molecular Docking, 3D-QSAR, and In Silico ADMETox Studies. Sci Rep. 2019;9(1):17096.
- Ramkissoon A, Seepersaud M, Maxwell A, Jayaraman J, Ramsubhag A. Isolation and Antibacterial Activity of Indole Alkaloids from Pseudomonas aeruginosa UWI-1. Molecules [Internet]. 2020; 25(16):1-14.
- Sonfack G, Fossi Tchinda C, Simo IK, Bitchagno GTM, Nganou BK, Çelik İ, et al. Saponin with antibacterial activity from the roots of Albizia adianthifolia. Natural Product Research. 2021;35(17):2831-2839.
- Tan Z, Deng J, Ye Q, Zhang Z. The Antibacterial Activity of Natural-derived Flavonoids. Curr Top Med Chem. 2022;22(12):1009-1019.
- Yuan G, Guan Y, Yi H, Lai S, Sun Y, Cao S. Antibacterial activity and mechanism of plant flavonoids to gram-positive bacteria predicted from their lipophilicities. Sci Rep. 2021;11(1):10471.
- Hussaini Y, Bello RY, Mustapha T. Preliminary phytochemical screening and GC-MS analysis of Anogeissus leiocarpus stem bark extract. The Pharma Innovation Journal. 2022;11(11):113-117.
- Mann A. Evaluation of antimicrobial activity of Anogeissus leiocarpus and Terminalia avicennioides against infectious diseases prevalent in hospital environments in Nigeria. Journal of Microbiology Research. 2012;2(1):6-10.
- Gara TY. In-vitro activity of methanolic extract of stem bark of Anogeisus leiocarpus (African birch) on Plasmodium falciparum. International Journal of Applied Biological Research. 2020;11(2): 69-78.
- Rathod M, Rakholiya K, Kaneria M, editors. Effects of Extraction Techniques on Phytochemical Analysis and Antioxidant Activity of Syzygium Cumini (L.) Skeels Seeds. Proceedings of the National Conference on Innovations in Biological Sciences (NCIBS) 2020. Available at http://dx.doi.org/10.2139/ssrn.3598090
- Edewor TI, Akpor OB, Owa SO. Determination of antibacterial activity, total phenolic, flavonoid and saponin contents in leaves of Anogeissus leiocarpus (DC.) Guill and Perr. J Coast Life Med. 2016;4(4):310-315.
- Timothy SY, Mashi FI, Helga BI, Galadima IH, Midala TAS. Phytochemical screening, antibacterial evaluation and in vitro spasmodic effect of the aqueous and ethanol leaf and bark extract of Anogeissus leiocarpus (DC) Guill. & Perr. Asian Journal of Pharmaceutical science & Technology. 2015;5(4): 208-302.
- Zumbes HJ, Belenu TO, Onwuliri FC. In vitro antibacterial activity of Anogeissus leiocarpus leaf extracts on some bacteria associated with diarrhea. International Journal of Natural and Applied Sciences. 2007;3(1):53-6.
- Abalaka ME, Daniyan SY, Mann A. Evaluation of the antimicrobial activities of two Ziziphus species (Ziziphus mauritiana L. and Ziziphus spinachristi L.) on some microbial pathogens. African Journal of Pharmacy and Pharmacology. 2010;4(4):135-144.
- Adejumobi JA, Ogundiya MO, Kolapo AL, Okunade MB. Phytochemical composition and in vitro antimicrobial activity of Anogeissus leiocarpus on some common oral pathogens. J Med Plants Res. 2008;2(8):193-199.
- Zinner SH, Mayer KH. 33 - Sulfonamides and Trimethoprim. In: Bennett JE, Dolin R, Blaser MJ, editors. Mandell, Douglas, and Bennett's Principles and Practice of Infectious Diseases (Eighth Edition). Philadelphia: W.B. Saunders; 2015. p. 410-8.e2.
- Pinto MEA, Araújo SG, Morais MI, Sa NP, Lima CM, Rosa CA, et al. Antifungal and antioxidant activity of fatty acid methyl esters from vegetable oils. An Acad Bras Cienc. 2017;89:1671-1681.
- Cavalca LB, Lahive CW, Gijsbers F, Pavan FR, Scheffers D-J, Deuss PJ. Benzenetriol-Derived Compounds against Citrus canker. Molecules [Internet]. 2021; 26(5):1-12.
- Sauvage E, Kerff F, Terrak M, Ayala JA, Charlier P. The penicillin-binding proteins: structure and role in peptidoglycan biosynthesis. FEMS Microbiol Rev. 2008;32(2):234-258.
- Tipper DJ. Mode of action of β-lactam antibiotics. Pharmacol Ther. 1985;27(1):1-35.
- European Bioinformatics Institute. Mechanism and Catalytic Site Atlas; Dihydrofolate reductase (bacterial) 2023 [cited 2023 22/3/2023]. Available from: https://www.ebi.ac.uk/thornton-srv/m-csa/entry/112/.
- Wróbel A, Maliszewski D, Baradyn M, Drozdowska D. Trimethoprim: An old antibacterial drug as a template to search for new targets. Synthesis, biological activity and molecular modeling study of novel trimethoprim analogs. Molecules. 2019;25(1):116.
- Kandasamy E, Kumar A, Rayasam GV, Perdih A, Bajpai U. Development of a one-pot assay for screening and identification of Mur pathway inhibitors in Mycobacterium tuberculosis. Sci Rep. 2016 10/13;6:35134.
- Petek M, Baebler Š, Kuzman D, Rotter A, Podlesek Z, Gruden K, et al. Revealing fosfomycin primary effect on Staphylococcus aureus transcriptome: modulation of cell envelope biosynthesis and phosphoenolpyruvate induced starvation. BMC Microbiol. 2010;10(1):1-12.
- Helgesen E, Sætre F, Skarstad K. Topoisomerase IV tracks behind the replication fork and the SeqA complex during DNA replication in Escherichia coli. Sci Rep. 2021;11(1):474.
- Pietsch F, Bergman JM, Brandis G, Marcusson LL, Zorzet A, Huseby DL, et al. Ciprofloxacin selects for RNA polymerase mutations with pleiotropic antibiotic resistance effects. The Journal of antimicrobial chemotherapy. 2017;72(10):175-184.
- Varshney A, Ansari Y, Zaidi N, Ahmad E, Badr G, Alam P, et al. Analysis of binding interaction between antibacterial ciprofloxacin and human serum albumin by spectroscopic techniques. Cell Biochem Biophys. 2014;70:93-101.
- Manganyi MC, Regnier T, Tchatchouang C-DK, Bezuidenhout CC, Ateba CN. Antibacterial activity of endophytic fungi isolated from Sceletium tortuosum L. (Kougoed). Annals of Microbiology. 2019;69(6):659-663.
- Vijayakumar K, Thirunanasambandham R. 5-Hydroxymethylfurfural inhibits Acinetobacter baumannii biofilms: An in vitro study. Arch Microbiol. 2021;203:673-682.
- Lu H, Tonge PJ. Inhibitors of FabI, an Enzyme Drug Target in the Bacterial Fatty Acid Biosynthesis Pathway. Accounts of Chemical Research. 2008;41(1):11-20.
- Wittke F, Vincent C, Chen J, Heller B, Kabler H, Overcash J, et al. Afabicin, a First-in-Class Antistaphylococcal Antibiotic, in the Treatment of Acute Bacterial Skin and Skin Structure Infections: Clinical Noninferiority to Vancomycin/Linezolid. Antimicrob Agents Chemother. 2020;64(10):e00250-20.
